"This web page was produced as an assignment for an undergraduate course at Davidson College."
My Favorite Yeast Genes
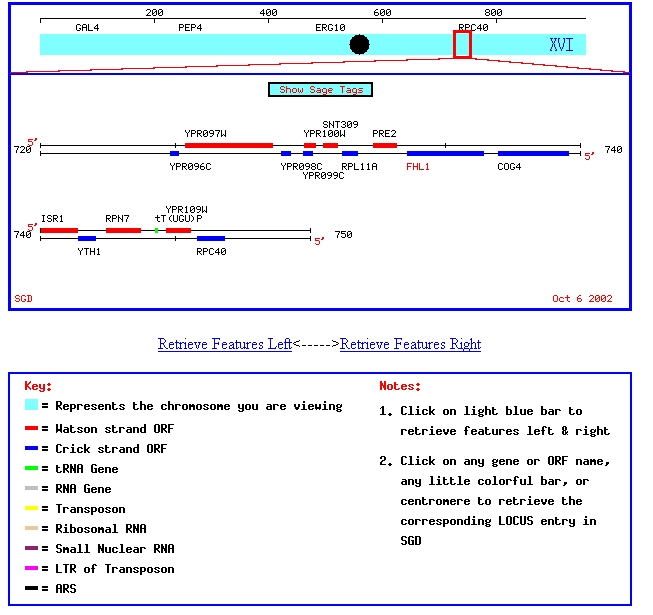
My favorite annotated yeast gene, FHL1, and my favorite non-annotated yeast gene, YPR098C, are located on chromosome XVI in Saccharomyces cerevisiae. This chromosomal map shows their positition on chromosome XVI:

Figure 1: Chromosomal map showing locations of FHL1 and YPR098C. Image from SGD, 2003 (http://genome-www4.stanford.edu/cgi-bin/SGD/ORFMAP/ORFmap?sgdid=S0006308)
My Favorite Annotated Gene
FHL1
The FHL1 yeast gene codes for a fork head protein involved in rRNA processing (SGD, 2003;http://genome-www4.stanford.edu/cgi-bin/SGD/GO/goAnnotation.pl?locus=FHL1). Therefore, using the definitions of Gene Ontology, the Biological Function and Process of the FHL1 yeast gene is to encode a transcription factor, Fhl1p, which regulates transcription of the RNA Polymerase III promoter which is involved in rRNA processing. Obviously, since the FHL1 gene encodes a transcription factor, it's cellular component is located in the nucleus. I was interested in this gene because the protein it encodes has a domain similar to the fork head DNA-binding domain found in the developmental fork head protein of Drosophila melanogaster and in the HNF-3 family of hepatocyte mammalian transcription factors and I wanted to learn more about its role in development in yeast (SGD, 2003; http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=FHL1).This gene is obviously very important to the Saccharomyces cerevisiae organism because a deletion of the gene leads to an inviable phenotype. In 1994, Hermann-Le et al. stated that null mutations caused a severe reduction in growth rate and a lower rRNA content that resulted from defective rRNA processing, but had no detectable effect on mRNA splicing. These results make sense due to the functional role of the FHL1 gene in yeast. Hermann-Le et al. also stated that FHL1 is a dosage-dependent suppressor and suggest that rRNA processing depends on a still-unidentified RNA polymerase III transcript.
One year later, Cherel and Thuriaux studied the interactions between Fhl1p and Ifh1p. They found that Ifh1p is essential for growth, but only in the presence of a functional Fhp1p protein. They also found that overexpression of Ifh1p partially compensates for the genetic inactivation of Fhl1p. They therefore concluded that their data suggest a direct interaction between the Fhl1p and Ifh1p proteins and are consistent with a model where Fhl1p is converted from a transcriptional repressor to an activator on binding of Ifh1p.
The fork head protein of Drosophila is a transcription factor that promotes terminal development. It contains neither homeodomains nor zinc-fingers characteristic of other transcription factors. Instead, it contains a distinct type of DNA-binding region called the fork head domain or winged helix. It contains around 100 amino acids and has since been identified in a number of transcription factors (including Drosophila FD1-5, mammalian HNF-3, human HTLF, yeast HCM1, yeast FHL1, etc.) involved in early developmental decisions of cell fates during embryogenesis. The fork head domain binds B-DNA as a monomer, but shows no similarity to previously identified DNA-binding motifs (Interpro, 2003; http://www.ebi.ac.uk/interpro/IEntry?ac=IPR001766#).
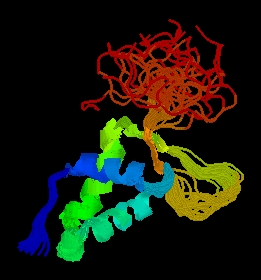
Figure 2: PDB picture of forkhead domain of FHL1 homolog, adipocyte- transcription factor freac-11 (s12) in humans. Cartoon structure view with different groups highlighted in different colors (PDB, 2003; http://www.rcsb.org/pdb/cgi/explore.cgi?job=graphics&pdbId=1d5v)
FHL1p has also been shown to interact with transcription initiation factors, DNA binding proteins, and protein tagging proteins. Predator analysis predicts that this protein should have mostly coiled coil structures and kyte-doolittle shows that this protein is not very hydrophic and therefore, not transmembrane domain. Although the yeast HCM1 gene also has a fork head domain, when I performed a BLAST2 protein analysis to compare its sequence to that of FHL1, I didn't observe much similarity between the two sequences.
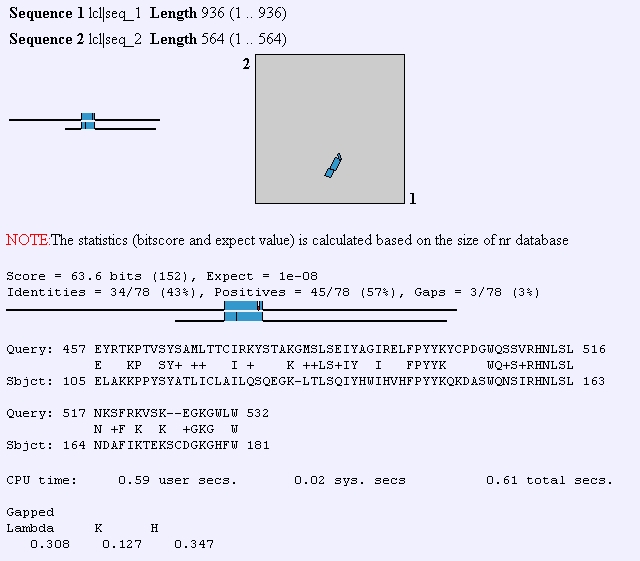
Figure 3: BLAST2 analysis of FHL1 protein sequence compared to HCM1 protein sequence. (http://www.ncbi.nlm.nih.gov/gorf/bl2.html)
The mammalian homologs link on the SGD web page stated that mouse myocyte nuclear factor, human transcription factor HFK2, and rat HNF-3/fork-head homolog-5 had the most similar protein sequences to the yeast gene FHL1.
The information above and the links below are pretty much a summary of the known information about the yeast gene FHL1 and its gene product FHL1p. Studies are ongoing on developmental proteins in yeast. In 1998, Zhu and Davis published a paper about Hcm1p and its regulation of SPC110, a gene that encodes the calmodulin-binding protein in the yeast spindle pole body (SGD, 2002; http://genome-www4.stanford.edu/cgi-bin/SGD/reference/reference.pl?refNo=25039). I will be interested to see future studies on FHL1p and its role in regulating rRNA processing and development.
Links to more information about the yeast gene FHL1:
Restriction Map : Type in FHL1
Protein Info: N and C terminal sequences, amino acid length, MW, pI, Hydropathicity
Fork Head Transcription Factor Info.
All you ever needed to know about FHL1 on one page! (http://genome-www4.stanford.edu/cgi-bin/SGD/singlepageformat?sgdid=S0006308)
My Favorite Non-Annotated Gene
YPR098C
Not much is known about the yeast gene YPR098C besides its nucleotide sequence. As seen above in the chromosomal map, it is located upstream of the FHL1 gene on chromosome XVI. Mutants lacking this gene are viable, so the YPR098C gene does not seem to be as vital to the Saccharomyces cerevisiae organism as the FHL1 gene (SGD, 2003; http://genome-www4.stanford.edu/cgi-bin/SGD/locus.pl?featureName=YPR098C). This is a very small hypothetical ORF as seen from the short amino acid sequence (108 amino acids):
MQAASPVILALTAPIALTTGPLSSLVVASVSGLTNLFWLLPWTHKVKEQRKNIAKKYTGS
ELEAKDAILRKEFGKSHGLSLLFNLSNVCGMLAYGVCLSGGLLRKIPK
Figure 4: Amino Acid Translation of yeast YPR098C gene. Data obtained
from SGD, 2003 (http://genome-www4.stanford.edu/cgi-bin/SGD/getSeq?seq=YPR098C&flankl=0&flankr=0&map=p3map)
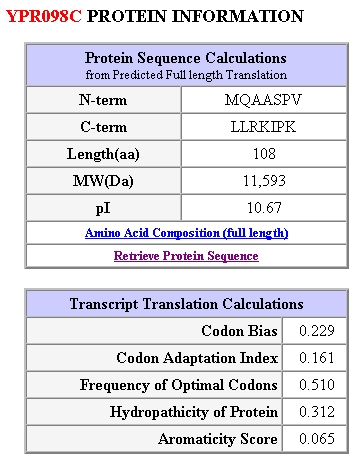
Figure 5: Protein Information for yeast gene YPR098C. Obtained from SGD, 2003 (http://genome-www4.stanford.edu/cgi-bin/SGD/protein/protein?sgdid=S0006302).
In an attempt to learn more about this gene I performed a BLASTn search with the DNA sequence. The first hit was the YPR098C gene itself, but the third hit was mouse connexin gene which is a gap junction protein expressed in neurons of the CNS. I thought this was interesting, but on a closer look, I realized that only 30 amino acids of 300 were similar. Therefore, the BLAST search gave me no additional information about this gene. A BLASTp search resulted in the identification of the yeast gene again as the top hit with the remaining hits not having enough similarity to be considered important. The BLASTp search did state that this hypothetical ORF was probably a membrane protein.
Therefore, I next performed a Kyte-Doolittle Analysis to analyze the protein sequence for hydrophic regions. Any section above the red line is a hydrophic region and possible transmembrane domain. There seem to be 3 to 5 possible transmembrane domains in this short sequence.
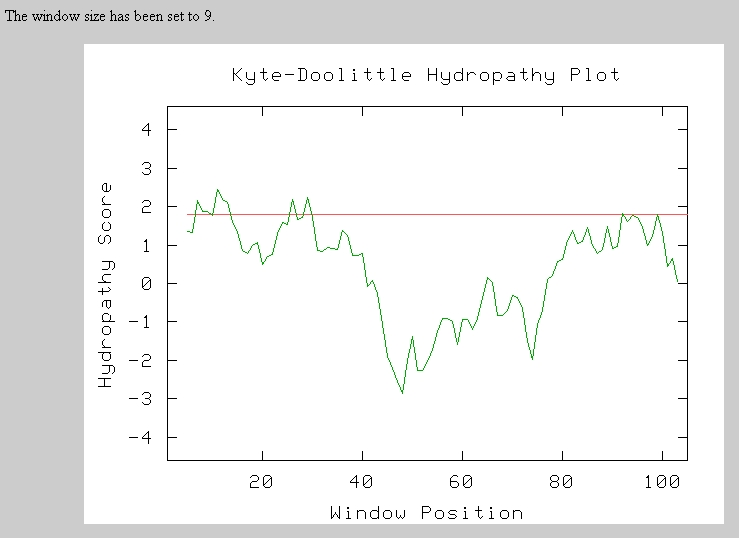
Predator Analysis predicted that this protein would be mostly random coils and alpha helices:
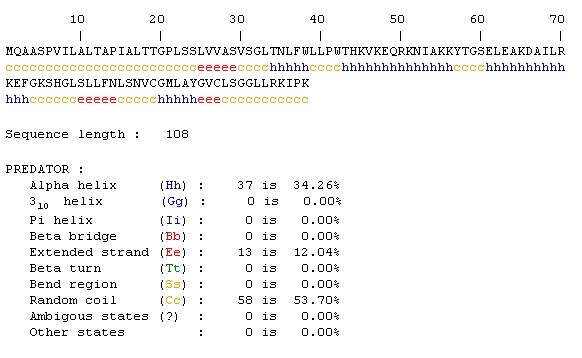
The Predator analysis didn't really provide that much additional information either. A conserved domain resulted in no hits, so I tried the mammalian homologs link from the SGD page. Again, the results initially looked promising, but only 10 amino acids were being compared.
In conclusion, I learned that the yeast gene YPR098C is 7486 nucleotides upstream of the yeast gene FHL1 that is a very small hypothetical ORF encoding a 108 amino acid product. Not much else is known about this gene besides its DNA sequence. There are no known homologs that I could compare it to and predictive webpages such as BLASTp and Predator didn't provide much information. A Kyte-Doolittle analysis did provide information to suggest that the protein produced by the YPR098C gene is a membrane protein, but sadly, that's about all I can conclude. Since a deletion of this gene leads to a viable phenotype, its role may not be important, but hopefully future research will be performed to elucidate the role of this gene in Saccharomyces cerevisiae.
SGD single page analysis of YPR098C gene
References:
Cherel I, Thuriaux P. 1995. The IFH1 gene product interacts with a fork head protein in Saccharomyces cerevisiae. Yeast 11(3):261-70.
Hermann-Le Denmat S, Werner M, Sentenac A, Thuriaux P. 1994. Suppression of yeast RNA polymerase III mutations by FHL1, a gene coding for a fork head protein involved in rRNA processing. Mol Cell Biol 14(5):2905-13.
Zhu G, Davis TN (1998) The fork head transcription factor Hcm1p participates in the regulation of SPC110, which encodes the calmodulin-binding protein in the yeast spindle pole body. Biochim Biophys Acta 1448(2):236-44.
Send comments, questions, and suggestions to: sahenry@davidson.edu