This web page was produced as an assignment for an undergraduate course at Davidson College.
My favorite annotated yeast gene, Gln3, and non-annotated yeast gene, Yen1, are both located on chromosome V in Saccharomyces cerevisiae. Below is a chromosomal features map showing their positions on chromosome V.
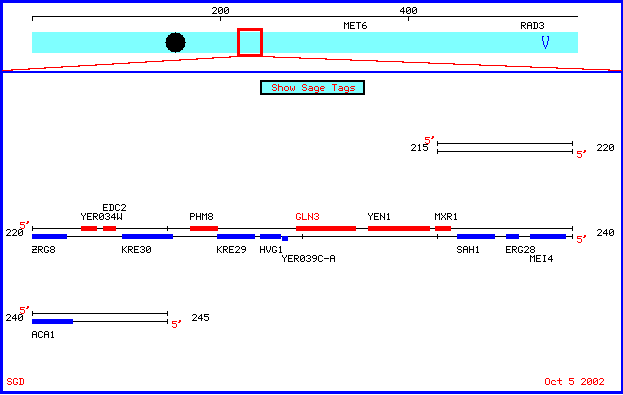
Figure 1: Chromosomal features map depicting locations of Gln3 and Yen1. Image obtained from SGD, 2003.
My Favorite Annotated Gene
The Gln3 gene encodes a transcription factor (gln3p) found in the nucleus and cytoplasm which plays a role in nitrogen catabolite repression (NCR). S. cerevisiae preferentially uses good nitrogen sources instead of bad ones "by repressing GATA-factor dependent transcription of the genes needed to transport and catabolize poor nitrogen sources"(Cox, et al., 2000, http://genome-www.stanford.edu/Saccharomyces/yeast00/abshtml/091.html). So when there are many nitrogen sources readily available, genes for proteins which help breakdown poor nitrogen sources are repressed. Genes sensitive to NCR produce two different sized transcripts. A longer mRNA is one product which is not sensitive to NCR and uses any nitrogen source, good or bad. The shorter mRNA from NCR sensitive genes is produced when Gln3p binds to a gene. The longer transcript is produced when Gln3p is not bound to GATAA site. The GATAA site is then used as an alternative transcriptional start site using GATAAs to replace normal TATAs. Alternatively, the shorter mRNA transcript is produced when Gln3p binds to NCR genes. Click here for gene sequence information.
It may sound simple but it is not! Gln3p is also regulated by cis-acting upstream activation sequence (UAS) which is found upstream of NCR-sensitive genes. The UAS's also contain the GATAA sequence. This prompted a group of researchers to determine if Gln3p bound to this UAS since Gln3p has a zinc finger motif (like most transcription factors) and GATAA sequence which is common to DNA binding factors. It was found that Gln3p does bind to this UAS in the presence of accessible nitrogen (Cunningham TS, et al., 1996, http://genome-www4.stanford.edu/cgi-bin/SGD/GO/goAnnotation.pl?locus=GLN3). Other research has shown that Gln3p can bind with Ure2p which consequently gets it excluded from the nucleus and into the cytoplasm. This hinders Gln3p from getting to the DNA and reaching the GATAA binding sites (Cox et al., 2000, http://genome-www.stanford.edu/Saccharomyces/yeast00/abshtml/091.html). Also Gln3p seems to be controlled by certain TOR proteins. TOR proteins are nutrient sensors and are regulated by nitrogen levels. One study shows that glutamine starvation caused nuclear localization of Gln3p (Gln3p is usually inhibited by TOR proteins). This follows logically that Gln3p would be nuclearized when the short transcript is needed (Crespo JL, et al. 2002,http://genome-www4.stanford.edu/cgi-bin/SGD/reference/geneinfo.pl?locus=GLN3 ) . Click here for the Gln3p protein sequence.
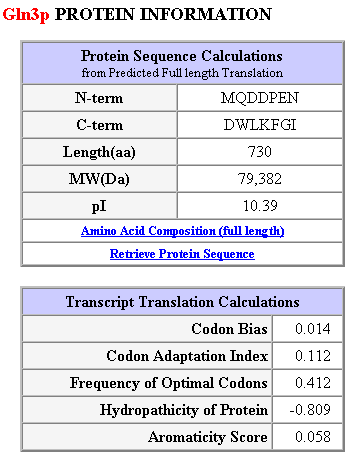
Figure 2: Gln3p protein information.
I couldn't download the image of Gln3p (found at this site http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) The protein has the GATA zinc finger conserved domain. I also used PREDATOR to predict the protein form. It is know it has a zinc finger and GATAA region. Below shows that Gln3p had alpha helices and many random coils.

Figure 3: Predator results of Gln3p protein sequence.
Mutant Gln3p yeast are viable, but do not grow well on poor nitrogen sources. As expected the longer mRNA transcript predominates in Gln3p mutants.
Gln3p is not highly conserved in many species. Fungus, chicken, and mice had high homology with the yeast Gln3p, but not humans. To view homology information, click here. This makes sense because humans do not use nitrogen as major energy source and would most likely have alternative pathways for nitrogen metabolism when needed. In other words, utilization of nitrogen is not a limiting factor in human growth.
My Favorite Non-Annotated Yeast Gene
YEN1 is located 474 nucleotides downstream from Gln3p on chromosome V. The molecular function and biological process of YEN1 are unknown. The gene sequence, however, is known and can be viewed by clicking here (along with the protein ORF translation). The YEN1 mutant is viable. This however does not rule out that it has an important function, but may be a redundant gene.
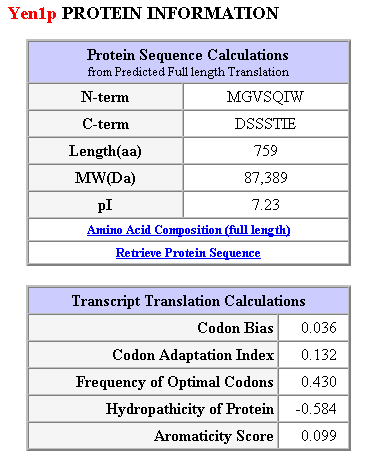
Figure 4: General protein information for YEN1.
Using PREDATOR I can make some predictions on the shape and function of the protein.

The protein looks a lot like Gln3p with random coils and alpha helices, except here there are more alpha helixes.
Using Blastp I can locate other organism that have protein sequences like that of YEN1 which may reveal something about its cellular role. The hits that came up related YEN1 protein sequence to DNA repair proteins and exonucleases in fungi. The bit scores were somewhat low but the E-value was low as well. To view the results click here.
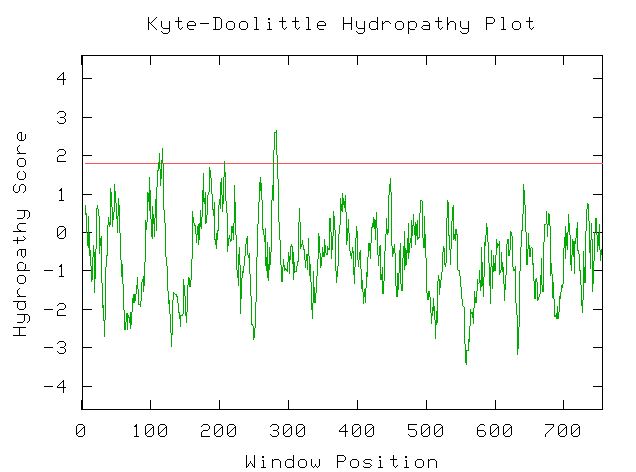
A Kyte-Doolittle Hydropathy plot with the amino acid sequence illustrates that this protein is most likely not an integral membrane protein.
YEN1 has a few conserved domains. It has a XPG I-region, a Xeroderma pigmentosum G I-region (domain in nucleases), and a Xeroderma pigmentosum GN-region (domain in nucleases). This supports the Blastp hits of fungi exonucleases with YEN1. (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi)
References
Cox, et al. "S. cerevisiae GATA sequences function as TATA elements during nitrogen catabolite repression because Gln3p is excluded from the nucleus." 2000. http://genome-www.stanford.edu/Saccharomyces/yeast00/abshtml/091.html
Crespo JL, et al. (2002) The TOR-controlled transcription activators
GLN3, RTG1, and RTG3 are regulated in response to intracellular levels of
glutamine. Proc Natl Acad Sci U S A 99(10):6784-9. http://genome-www4.stanford.edu/cgi-bin/SGD/reference/geneinfo.pl?locus=GLN3
Cunningham, TS, et al. "Gln3p Is Capable of Binding UAS Elements and Activating Transcription in Saccharomyces cerevisiae." 1996.
SGD database: http://genome-www.stanford.edu/Saccharomyces/