*This web page was created as an assignment for an undergraduate course at Davidson College*
TKL1 Protein information: I investigated the protein function of TKL1. TKL1 is a yeast gene known to function as a transketolase enzyme. This enzyme functions as part of the pentose phosphate pathway. For more information on the role of TKL1 please see my second web assignment (YFYG). For information on the expression patterns of TKL1 please see my third web assignment (YFYE).
MIPS information:
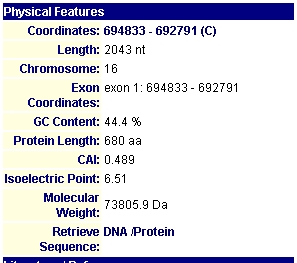
By searching the MIPS database I found information on the protein encoded by TKL1. The figure below indicates TKL1's pI and its molecular weight. This figure also shows the number of amino acids in the TKL1 protein.

Figure 1: MIPs information on the physical features of the TKL1 protein and gene. The protein's length is 680 amino acids. ITs molecular weight is 73805.9Daltons and it has an isoelectric point of 6.51. (http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YPR074)
The next figure shows that TKL1 has no hits for possible protein protein interactions. It does however have possible hits for a protein complex with TKL2.
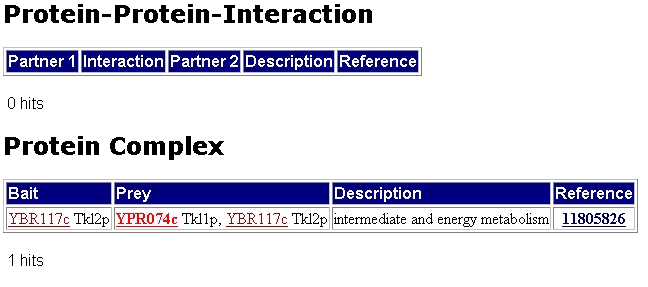

Figure 2: MIPs information on possible protein-protein interaction and possible protein complex of TKL1. Only one hit is found which indicates that TKL1 may form a complex with TKL2.(http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YPR074c)
The MIPs database indicates that TKL1 and TKL2 are share a strong similarity. The ratio of TKL1/TKL2 in a cell varies depending on the physiological state of the cell. TKL2 has different thermostability than that of TKL1. MIPS also indicates that TKL2 functions as a dimer. This database claims that the TKL1p/TKL2p is required for the biosynthesis of erythrose-4-phosphate. This indicates that TKL1 may function as a dimer with TKL2 in order to catalyze reactions in the pentose phosphate pathway. (http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YPR074c)
PDB:
After searching MIPS I then searched PDB for further information on the TKL1 protein. This database lead me to several pubmed articles indicating the role of the TKL1 protein in catalytic activities.
The first article identified catalytically important residues in yeast transketolase. This article indicates that specific amino acid residues are required for the yeast transketolase to function properly. To see the abstract to this article click here: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=9398292.
The second entry was for a pubmed article indicating the substrate binding domains for TKL1. This article describes the domains on yeast transketolase in which its substrate, erythrose -4- phosphate binds. To see the abstract of this article click here: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=8999873.
The third article discusses the specificity of binding between transketolase and its coenzyme thiamine diphosphate To view the abstract from this article click here: http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Abstract&list_uids=8144674.
ScanProsite:
In this database I entered the PDB ID of 1NGS. I then scanned prosite for possible posttranslational modification domains. This site listed a glycosylation site, a cAMP dependent cGMP dependent protein kinase phosphorylation site, a Casein kinase 2 phosphorylation site and an N-myristolation site. All of these sites contained high probability warnings. Signature transketolase 1 and 2 sites were found. In their documentation of TKL1 prosite lists the information found in figure 4.
 .
.
Figure 4: TKL1 documentation from prosite database. Indicates that TKL1 acts as a dimer and requires thiamine pyrophosphate as a cofactor. The final paragraph also indicates the two regions of TKL1 that are TK signature patterns. (http://us.expasy.org/cgi-bin/nicedoc.pl?PDOC00635).
Database of Interacting Proteins (DIP):
In this database I searched for the proteins that have been found to interact with TKL1. Upon this search I found the graph seen in figure 5. This graph indicates that TKL1 interacts with TKL2. TKL1 is the large red dot and TKL2 is its interacting protein, the smaller orange dot. Based on the legend the red line between the two dots indicates that these results are unverified from high throughput interaction screens.
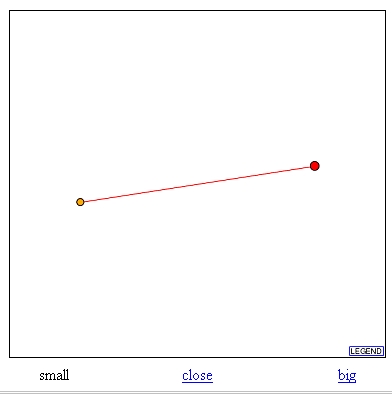
Figure 5: Graph from DIP database showing interaction between TKL1 and TKL2. TKl1 is red dot and TKL2 is the orange dot. (http://dip.doe-mbi.ucla.edu/dip/DIPview.cgi?PK=757).
Expasy 2D:
In this database I searched for TKL1. I then computed the theoretical molecular weight and pI. I got a molecular weight of 73674.45daltons and a theoretical pI of 6.50. These numbers closely matched the entries I received in the MIPS database. (http://ca.expasy.org/cgi-bin/pi_tool1?P23254@noft@).
I then examined the 2d gel for Saccharomyces cerevisiae. Figure 6 shows a 2d gel with the Saccharomyces cerevisiae proteins. The yellow box surrounds a region of the gel in which TKL1 proteins should be found. The green and red dots indicate the TKL1 proteins identified through sequence analysis in the gel.
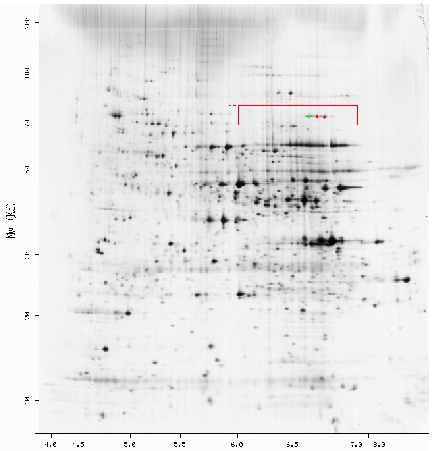
Figure 6: A 2D gel of Saccharomyces cerevisiae proteins. The red box indicates the possible location of the TKL1 protein. The green and red dots indicate the identification of TKL1 proteins. (http://ca.expasy.org/cgi-bin/ch2d-compute-map?YEAST,P23254,2D-000IUQ).
Databases searched:
I also searched for TKL1 in the pdf files from the Benno Scwikowski paper found at the following web site: http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/chapter6/deluxe.html. I found no hits for TKL1 in these pdfs. I also looked under Y2H results and found no information for TKL1.
Conclusions:
The information in the searched databases verify the fact that TKL1 is a transketolase that functions in the pentose phosphate pathway of saccharomyces cerevisiae. These databases indicated that the TKL1 protein often functions as a dimer and forms a complex with the similar protein TKL2. TKL1 has a molecular weight of 73805.9Daltons and a pI of 6.5.
Possible Further Investigations:
While much is already known about the TKL1 yeast gene and its gene product, transketolase 1, there is a still many experiments that can be conducted to further characterize the yeast TKL1p. First the change in relative abundance of transketolase 1 in different cellular environments can be investigated. Using Chait et al.'s stable isotope method relative quantities of protein can be compared(Campbell, 2002). In this method proteins in different cell cultures in varying cellular environments are labeled with different stable isotopes. The proteins from each cell culture are then combined and run on a 2D gel. The tranksketolase spots would then be analyzed through MALDI-MS/MS which would separate the labeled transketolase enzymes. By comparing the height of peaks the relative abundance of the enzyme in each cellular environment can be determined. This experiment would offer an insight into the level of transketolase needed in varying cellular environment. Due to the fact that transketolase functions in the pentose phosphate pathway, this experiment would help indicate under what cellular conditions this pathway was utilized the most.
In order to further investigate this enzyme the enzyme assay methods developed
by Dovichi et al could be utilized (Campbell, 2002). Based on these methods
single molecules of transketolase could be isolated. Using an enzymatic assay
the reaction rate for each enzyme molecule could be found and compared to other
enzyme molecules. A difference in reaction rats between identical enzymes grown
in the same cellular conditions would indicate that transketolase underwent
some form of modification that lead to its varying reaction rates.
These are just a few examples of the many experiments that could be used to further investigate the role of the TKL1 protein in yeast.
References:
Campbell, M., Heyer L. Dicovering Genomics Proteomics & Bioinformatics. Cold Spring Harbor Laboratory and Benjamin Cummings. San Francisco. 2003
Schwikowski, Benno, Uetz P., Fields S., A network of protein-protein interactions in yeast. Nature Biotechnology. 18: 1257-1261.