"This web page was produced as an assignment for an undergraduate course at Davidson College"
MIF2 and YKL091C
PROTEIN ANALYSIS

"This web page was produced as an assignment for an undergraduate course at Davidson College"
MIF2 and YKL091C
PROTEIN ANALYSIS

In the previous webpage assignment, the transcriptome of the yeast genome was analyzed using microarrays in order to determine the expression patterns of the genes MIF2 and YKL091C under varying conditions. The purpose of this webpage, however, is to analyze the proteome of the yeast S. cerevisiae in order to gain greater insight into the functions of the proteins encoded by these two genes. As stated in the previous assignments, the annotated MIF2 gene encodes for a protein essential to the mitotic process in S. cerevisiae. Specifically, MIF2 encodes for a centromeric protein involved in mitotic spindle structural assembly. The unannotated YKL091C gene is hypothesized to encode for a PITP, phosphatidylinositol transfer-like protein, involved in modulation of signal transduction and cross-membrane transportation.
DIP
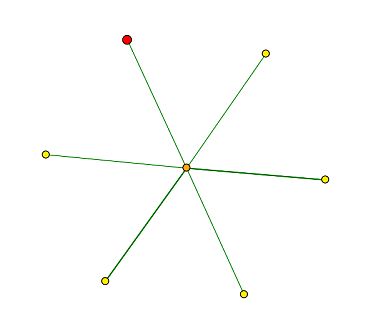
Figure 1: Screen capture of the graph generated by DIP for MIF2, which depicts the protein-protein interactions for the MIF2 encoded protein. The MIF2 protein is signified by the red node in the top left corner of the figure. MIF2 is shown to interact only with the central node, which represents the MCM21 protein. According to the Saccharomyces Genome Database, MCM21 is involved in chromsome segregation, and it also localizes in the condensed nuclear chromosome kinetechore.
The DIP database is useful for determining protein-protein interactions. It presents the interaction results as a graph in which the desired protein is signified by a red node, and the nodes connected to it signify the proteins that are suspected to interact with it. The greater the evidence to support the interaction between the two proteins, the thicker the line between the two nodes. By clicking on the node, one can ascertain the identity of the protein and other related information.
Benno Figure 1
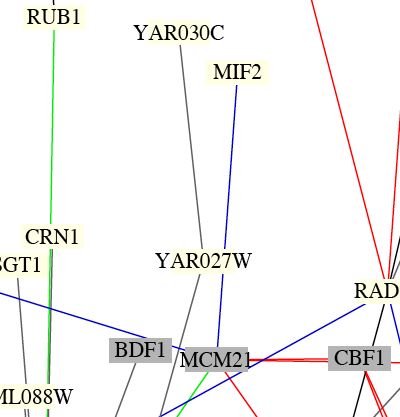
Figure 2: Screen capture depicting the position of MIF2 protein within the Benno Schwikowski figure, which depicts 2,709 interactions among 2,039 proteins. According to this Benno figure, the MIF2 protein interacts with the protein encoded by MCM21. The interaction displayed in this figure corroborates the information supplied by the DIP databse, which also indicates an interaction between MIF2 and MCM21. The blue line connecting the two signifies identical locations, but differences in function. Both the MIF2 and MCM21 proteins localize at the chromosome kinetochore.
Benno Schwikowski was a computer scientist, who collaborated with Stan Fields and Peter Uetz, in order to generate an interaction map for yeast protein. After compiling data concerning the 2,709 interactions among 2,309 proteins, Benno was able to generate complex diagrams of these interactions, of which a small portion is depicted in the figure above.
MIPS

Figure 3: Screen capture of the MIPS list of 8 interactions for MIF2 of which three are actual protein-protein interactions that are signified by the interaction label ">physical<". The proteins with which the MIF2 protein interacted included CBF1, MCM21 and CHL4. CBF1 is a centromere binding factor that is involved in chromosomal segregation. MCM21 is an outer kinetechore protein that plays a role in chromosomal segregation. Not surprisingly, CHL4 is also a outer kinetechore protein that is involved in chromosomal segregation.
MIPS is yet another database, which provides information regarding the genome of the yeast S. cerevisiae. This database also supplies information regarding the interactions of the proteins encoded by the genes. The protein interaction information for a gene is listed under the section entitled: PPI and Complex Viewer.
Prowl
The prowl search only served to confirm that the MIF2 protein is a centromeric protein that does in fact function in chromosome segregation and in the maintainence of spindle integrity. One of the Prowl hits resulting from my MIF2 search was my unannotated gene YKL091C.
Y2H
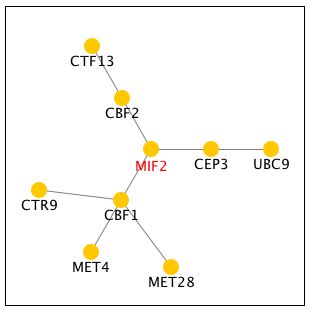
Figure 5: Screen capture of the genetic interactions of MIF2 with CBF1, CBF2, and CEP3. CEP3 is a component of the CBF3 kinetechore protein complex. CBF1 is centromere binding factor that is involved in mitotic segregation. CBF2 is another centromere binding factor, which is also a component of the CBF3 kinetechore protein complex.
The yeast-two hybrid method involves using the protein of interest as "bait" in order to determine the "prey" proteins, which physically interact with it.
Experiment 1 design
From the information uncovered up to this point concerning MIF2, it is clear that the protein for which MIF2 encodes is a centromeric protein. The protein interaction information supplied by the various databases cited above also indicates that the MIF2 protein solely interacts with proteins localized in the chromosomal kinetchore and involved in chromosomal segregation. In order to confirm MIF2's role in the mitotic process, a knockout could be generated in which the MIF2 gene has been deleted. This would undoubtedly result in an inviable phenotype because the cells would be prevented from completion of mitosis due to the lack of MIF2 protein. This outcome would result because the MIF2 protein is essential to the maintainence of the structural integrity of the spindle assembly prior to the anaphase spindle elongation step of the mitotic cycle. Another potential experiment might employ the yeast-two hybrid method, in which the bait protein (MIF2) is placed within the single MIF2 mutant cell during three stages in the mitotic cycle: prophase, metaphase and anaphase. This series of experiments would reveal that the MIF2 protein experiences no protein-protein interaction during metaphase or prophase; however, experiences an abundance of interaction during the initiation of anaphase during which it is essential to spindle integrity.
_________________________________________________________________________________________________
DIP
The DIP database did not have a operational protein interaction graph for the YKL091C protein.
Benno Figure 1
The protein encoded by the unannotated YKL091C gene was not one of the 2,039 proteins included in the diagram generated by Benno Schwikowski.
MIPS

Figure 5: Screen capture of the MIPS list of protein-protein interactions in which YKL091C is known to be involved. The MIPS database had no information with regard to protein interactions involving the protein encoded by YKL091C.
YKL091C PPI and Complex Viewer
Prowl
The prowl search did not yield any new information regarding the unannotated gene YKL091C or its corresponding protein.
Y2H

Figure 6: Screen capture of the known interactions of YKL091C. The Y2H database provided no information regarding known or even projected interactions for the YKL091C protein.
Experiment 2 Design
One potential method for discerning the function of the YKL091C gene and the function of the protein for which it encodes is with a good old fashioned knock-out. By knocking out a gene either by deletion or severe disruption, one can determine its function due to the developmental complications that arise for lack of the gene and its corresponding protein. I would use an experimental method similar to that outlined in the Giaever paper. The researchers involved in this experiment generated gene-deleted mutants in order to gain greater insight into the functions of both known and unknown genes in the yeast S. cerevisiae (2002). Groups of yeast cells with the YKL091C gene deletion could also be subjected to a number of variable conditions, and the results could then be contrasted with groups of yeast cells reared in the same conditions without the deletion. Since I hypothesized that YKL091C is an aerobic gene that encodes for a PITP, I would want to contrast the performance of a normal group of yeast cells with that of gene-deleted mutant cells in a high oxygen environment. Because the mutant lacks the YKL091C, I would expect the mutants to exhibit slower growth rates resulting from diminished efficiency in the areas of signal transduction and membrane trafficking. However, I would also conjecture that the phenotype would still be viable. In order to gain greater insight into potential protein-protein interaction involving the YKL091C, I would also propose the execution of an experiment that utilizes the yeast-two hybrid method in which the YKL091C protein is used as the bait. When the bait is fused with the DNA binding domain and placed within a cell, any interaction with a prey protein will result in the transcription of a reporter gene. This experiment would be a good first step towards a better understanding of protein-protein interaction involving YKL091C.
Links:
References:
Campbell, AM and Heyer, LJ. Genomics, Proteomics, & Bioinformatics
2003. San Francisco, CA: Pearson Education, Inc.
Database of Interacting Proteins (DIP), 2001, <http://dip.doe-mbi.ucla.edu/dip/Search.cgi?SM=3>
MIPS Comprehensive Yeast Genome Database (CYGD), 2002, <http://mips.gsf.de/genre/proj/yeast/index.jsp>
Saccharomyces Genome Database (SGD), 2002, <http://www.yeastgenome.org/>
Prowl, 2000, <http://129.85.19.192/prowl/proteininfo.html>
Yeast Interaction Database, 2002, <http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastList?modeIn=List
"MIF2/YKL089W." Saccharomyces Genome Database. Last
Update: 2003. <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?sgdid=S0001572>
"YKL091C." Saccharomyces Genome Database. Last Update: 2003. <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=YKL091C>
Giaever G, et al. (2002) Functional profiling of the Saccharomyces cerevisiae genome. Nature 418(6896):387-91
© Copyright 2003 Department of Biology, Davidson College, Davidson,
NC 28035
Send comments, questions, and suggestions to: dacheuy@davidson.edu