This web page was produced as an assignment for an undergraduate course at Davidson College
*My Favorite Yeast Genes: Fis1 and YIL067C*
Introduction
This web page explores two genes in the same chromosomal region of the organism S. cerevisiae, commonly known as Baker's yeast. One of the genes, Fis1, is an annotated gene with a known function, while YIL067C is a yet non-annotated gene. I will explore characteristics of each gene using various internet resources, eventually predicting a function for YIL067C.
Where in the S.cerevisiae genome are Fis1 and YIL064W located?
Fis1 and YIL067C are both located on yeast chromosome 9. The
chromosomal coordinates for Fis1 are
241772 - 241305 and the chromosomal coordinates for the ORF YIL067C are 237757
- 235721.
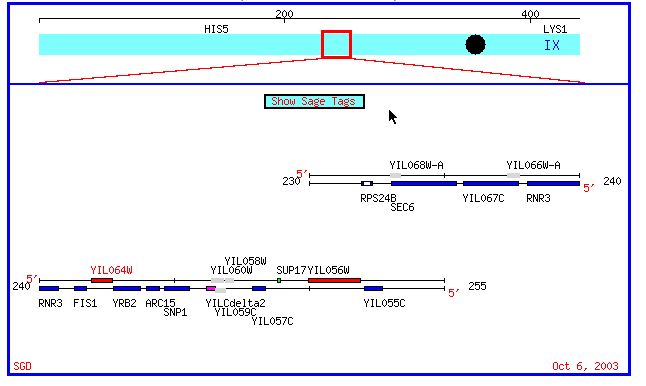
Figure 1: Chromosomal map of yeast chromosome 9 region containing both Fis1 and YIL067C. (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/ORFMAP/ORFmap?sgdid=S0001327>).
Fis1
Fis1 encodes a protein that is 155 amino acids long and weighs 17.7 kDa (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/protein/protein?sgdid=S0001327>). Other names for Fis1 include YIL065c and MDV2 (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=Fis1>).
Click here for Fis1's genomic sequence.
Click here to access Fis1's Entrez protein citation.
Fis1's MIPS citation.
Biological Process (Why?)
Some databases state that the biological process of Fis1 is unknown (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=Fis1>). However, fusion and fission together regulate the morphology of mitochondria. It seems reasonable to conclude that mitochondrial morphology, which is highly directed, plays an important role in overall mitochondrial function, which is to store energy from food in ATP. Therefore, I conclude that Fis1's Biological Function is energy storage and energy distribution to cell structures. It has been suggested that fusion and fission events help direct the speread of mitochondrial membrane potential to energy starved regions of the cell, further supporting my conclusion (Skulachev, 1990).
Molecular Function (What?)
Fis1 protein is essential to mitochondrial fission. There are other yeast proteins that are essential to mitochondrial division as well; two of these are Dnm1p and Mdv1p. Fis1p is necessary in order to localize Dnm1p and Mdv1p to the outer mitochondrial membrane; one review article called it a sort of 'assembly receptor' (Shaw 2002). By facilitating assembly of these complexes, Fis1 regulates the distribution and location of mitochondrial outer membrane fission events. More specifically, it has been suggested that Fis1p initially only interacts with Dnm1p, which in turn interacts with Mdv1p. Througout the membrane remodeling process, Mdv1p may move to interact with Fis1p in one of two fashions, facilitating the actual fission step. These hypotheses are depicted in the diagram below.
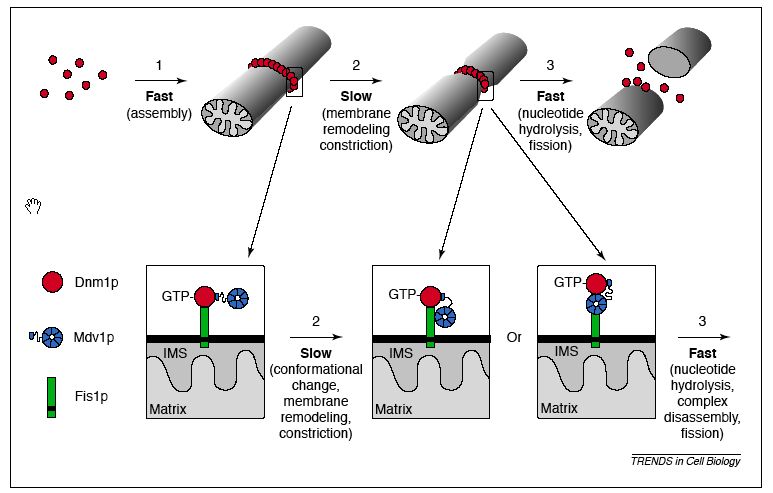
Figure 2: Proposed molecular model of mitochondrial fission: interactions between Dnm1p, Mdv1p, and Fis1p. Note: IMS stands for the intermembrane space of mitochondria. From: Shaw & Nunnari, 2002. Permission requested.
Cellular Component (Where?)
Fis1 protein localizes to the outer membrane of mitochondria, as depicted below. Its amino terminus points into the cytoplasm of the cell (outside of the mitochondria), and its C terminus points into the intramembraneous space of the mitochondria (Mozdy, 2000).

Figure 3: Schematic of Fis1 gene orientation with respect to location of Fis1p on the outer mitochondrial membrane. From: Shaw & Nunnari, 2002. Permission requested.
Protien information from various web resources
I used the Kyte-Doolittle hydropathy plot to find probable transmembrane regions in Fis1p. I expected to see some transmembrane regions (peaks above 1.8), so this figure complements the information I found in other databases. This figure was obtained with the window size set at 9, and when I tried the plot with the window size set at 11, these same peaks were present, giving further confirmation to Fis1's presence on the outer mitochondrial membrane.
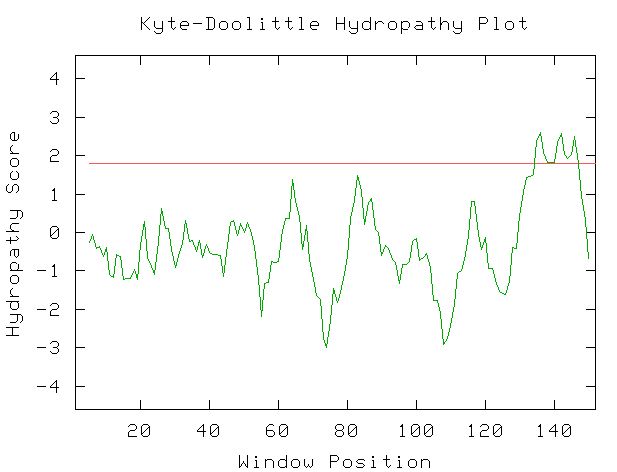
Figure 4: Kyte-Doolittle Hydropathy plot created by entering the amino acid sequence of Fis1p. Window size set at 9. Red line is at 1.8 hydropathy score.
When I searched PDB for Fis1, the only structure I found was named Rsgi Ruh-001 and came from mouse cDNA. The PDB citation stated that it resembled FIS1p and had an unknown function. An alignment of these two proteins from SGD showed 28% identity and 41% similarity between the two protein sequences (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/protein/get3d?locus=YIL065C&pdb=1iyg_A&align=1>).
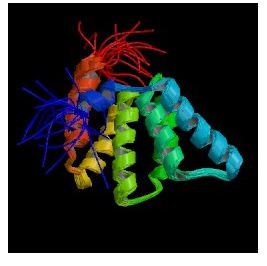
Figure 5: Still cartoon image of Rsgi Ruh-001 (PDB, 2003; <http://www.rcsb.org/pdb/cgi/explore.cgi?job=graphics&pdbId=1IYG&page=&pid=164711065585003>).
I searched PREDATOR with the amino acid sequence of Fis1 to try and reveal some of Fis1's secondary structure. The results I found are pictured below. The blue lines indicate alpha helices, and there appear to be six distinct regions of alpha helices. This correlates with the PDB image pictured in Figure 5, as there are six alpha helices (the spirals) pictured. The red lines indicate extended strands, and there appear to be two distinct groups of extended strands in both Figures 6 and 5. This gives further support to mouse Rsgi Ruh-001 having a very similar structure to Fis1.
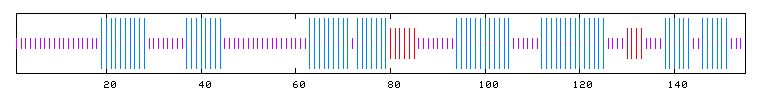
Figure 6: PREDATOR results for FIS1p. Purple lines denote random coils, blue lines indicate alpha helices, and red lines indicate extended strands (PREDATOR, 2003; <http://npsa-pbil.ibcp.fr/cgi-bin/secpred_preda.pl>).
I performed a BLASTp with Fis1's amino acid sequence. The results showed that two proteins have highly significant homology to Fis1p and that many other proteins have a notable sequence alignment. The top two scores were for a hypothetical neurospora protein and a putative mitochondrial fission protein in Tuber borchii (whitish truffle). Entrez protein links to these two proteins can be accessed by clicking on their names. Other notable alignments came from S. pombe, Anopheles gambiae, and the Norwegian rat.
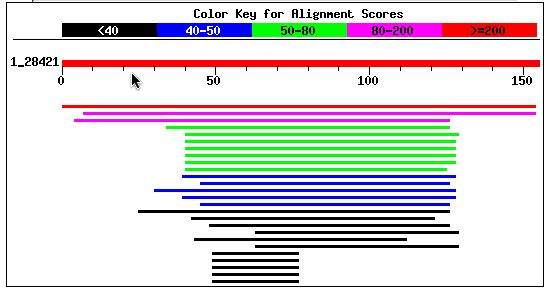
Figure 7: BLASTp results using Fis1's amino acid sequence (NCBI BLASTp, 2003; <http://www.ncbi.nlm.nih.gov/blast/>).
An NCBI Conserved Domain search using the Fis1 amino acid sequence as a query yielded no results.
What happens to yeast with mutant Fis1?
When Fis1 is mutated, yeast are still viable. However, their mitochondria are not capable of undergoing fission, and because mitochondria can still fuse, mitochondrial membranes accumulate in nets (Shaw, 2002). In Fis1 mutants with normal Dnm1 and Mdv1, complexes of these two proteins did not assemble properly and were present in large, undistributed clumps on the outer mitochondrial membrane as depicted below (Shaw, 2002).
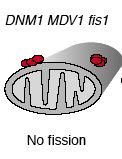
Figure 8: Diagrammatic representation of mitochondria from yeast with Fis1 mutated. From: Shaw & Nunnari, 2002. Permission requested.
Mutations in Fis1 have no effect on mitochondrial membrane fusion events.
What about Fis1 in other organisms?
Fis1 has been conserved in many organisms; it has homologs in eukaryotes such as worms, mammals and plants (Shaw, 2002). The NCBI BLASTp search performed above also confirmed that Fis1 has homologs in other organisms. In a recent experiment using a mammalian homolog, researchers found that the C-terminal structure of mammalian Fis1 was required for mitochondrial localization and that the N-terminal region was required for mitochondrial fission. When mammalian Fis1 was overexpressed, elevated fragmenting of mitochondria was seen, and when anti-mammalian Fis1 was applied to cells, elongated, non-divided mitochondria were seen (Yoon et al., 2003). These results concur with results seen in yeast experiments discussed previously, indicating that Fis1's function has been highly conserved.
YIL067C
YIL067C is 678 amino acids in length and has a hypothesized molecular weight of 75.5 kDa (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/protein/protein?sgdid=S0001329>)
Click here for the Entrez Protein link to YIL067C.
Click here for YIL067C's genomic sequence.
I performed a BLASTp with YIL067C's amino acid sequence and found that it shared significant amino acid similarity with two other potential genes. The first gene (second red line) was a hypothetical protein from the fungus Neurospora crassa that shared 47% positives and 30% identities with YIL067C. The expcet value for this similarity was 6e-50 (NCBI's BLASTp, 2003; <http://www.ncbi.nlm.nih.gov/blast/Blast.cgi>). The Entrez protein link to this hypothetical protein is can be accessed by clicking here. The second gene (magenta line) was also a hypothetical protein from Neurospora. It shared 44% positives and 30% identities with YIL067C. The expect value for this was 2e-34 (NCBI's BLASTp, 2003; <http://www.ncbi.nlm.nih.gov/blast/Blast.cgi>). The Entrez protein link to this probable protein can be accessed by clicking here. Each of these citations has the same amino acid sequence, indicating that each of these hits was for the same gene. Regardless, this information was not particularly helpful since the gene has not been annotated.
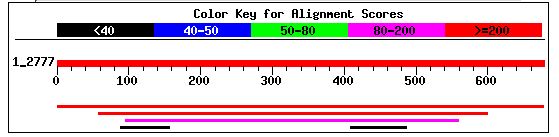
Figure 9: NCBI BLASTp hits after using YIL067C amino acid sequence as a query (NCBI BLAST, 2003 <http://www.ncbi.nlm.nih.gov/blast/>).
When I performed an NCBI conserved domain search with YIL067C's amino acid sequence, I found no hits! (Result pictured below).
Figure 10: NCBI Conserved Domain results using YIL067C's amino acid sequence as a query (NCBI, 2003; <http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi>.
However, I also decided to try a CD search with the amino acid sequence of the hypothetical protein homolog I found in BLASTp. When I used the sequence of the homolog, I found a conserved LCCL domain, as pictured below. No explicit function for the LCCL domain has been proposed yet, but it consists of a pattern of alpha helices and beta strands. Mutations in LCCL domains in certain human proteins have been aassociated with deafness (Grabski et al., 2003). Problematic mutations in LCCL domains affect protein folding, suggesting that the tertiary structure of this region is essential for protein function.
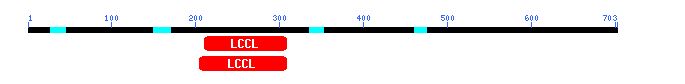
Figure 11: CD search using potential protein and Fis1 homolog from Neurospora (NCBI's CD, 2003; <http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi>).
I used the Kyte-Doolittle Hydryopathy plot to see if YIL067C contains any probable transmembrane domains. The figure below shows eight or nine probable transmembrane domains (peaks above 1.8). This figure was created using the potential amino acid sequence of YIL067C and the window size was set at 9 amino acids. This confirms comments found in databases pitting YIL067C as a putative intermembrane protein (Swissprot, 2003; <http://us.expasy.org/cgi-bin/niceprot.pl?P40514>). Additionally, under 'features', YIL067C's Swissprot citation noted 9 probable transmembrane domains.
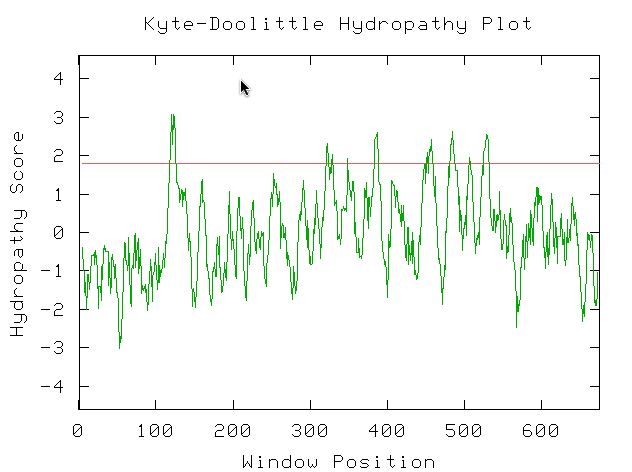
Figure 12: NCBI Conserved Domain results using YIL067C's amino acid sequence as a query (NCBI, 2003; <http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi>.
I used PREDATOR to try and shed some light on YIL067C's secondary structure. I can see from the figure below that there are multiple alpha helices (blue lines) and extended sequences (red lines).
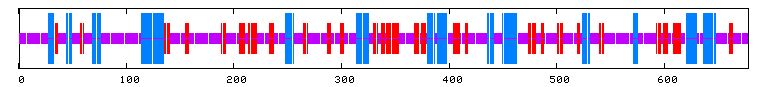
Figure 13: PREDATOR results for YIL067C. Purple lines denote random coils, blue lines indicate alpha helices, and red lines indicate extended strands (PREDATOR, 2003; <http://npsa-pbil.ibcp.fr/cgi-bin/secpred_preda.pl> .
Predicting a probable function for YIL067C is difficult considering its only homologs are probable proteins with unknown function and that it produced no hits in a search of conserved domains. I can predict with confidence that YIL067C is an intermembrane protein. The presence of the LCCL domain in the fungal homolog suggests that the same area of YIL067 may be particularly important in the way it folds, but that is about all I can deduce.
References
Grabski R., et al. 2003. Mutations in COCH that result in non-syndromic autosomal dominant deafness (DFNA9) affect matrix deposition of cochlin.Hum Genet. 113(5): 406-16.
Jakobs, S., et al. 2003. Spatial and temporal dynamics of budding yeast mitochondria lacking the division component Fis1p. J Cell Sci. 116(Pt.10): 2005-14.
Liepinsh, E., et al. 2001. NMR structure of the LCCL domain and implications for DFNA9 deafness disorder.EMBO J. 20(19):5347-53.
Mozdy, A.D., et al. 2000. Dnm1p GTPase-mediated mitochondrial fission is a multi-step process requiring the novel integral membrane component Fis1p. J Cell Biol. 151(2): 367-80.
Shaw, J.M. & Nunnari, J. 2002. Mitochondrial dynamics and division in budding yeast. TRENDS in Cell Biol. 12(4): 178-184.
Skulachev, V.P. 1990. Power transmission along biological membranes. J. Membr. Biol. 114; 97-112.
Yoon, Y., et al. 2003. the mitochondrial protein hFis1regulates mitochondrial fusion in mammalian cells through an interaction with dynamin-like protein DLP1. Mol Cell Biol 23(15):. 5409-20.
Questions, Comments? E-mail Sara