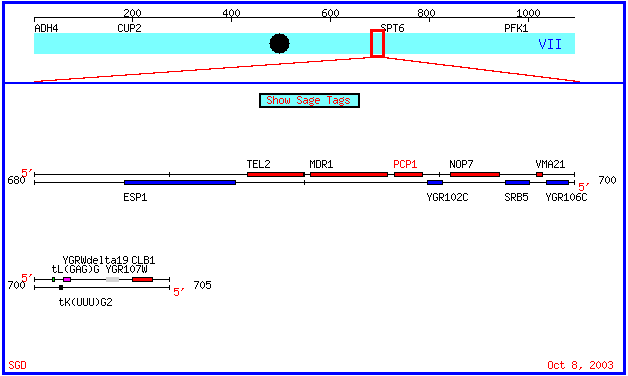
PCP1(also referred to by the systematic name YGR101W) and YGR102C are two genes of the organism Saccharomyces cerevisiae, commonly known as Baker's yeast. They are located within a small region in chromosome VII. While the function of PCP1 is well known, the nearby gene YGR102C is a hypothetical ORF that has not yet been annotated. The chromosomal map below, from the Stanford Saccharomyces Genome Database, shows the locations of the two genes. The chromosomal coordinates for PCP1 are 693364 to 694404 and the chromosomal coordinates for the YGR102C are 695136 to 694585.

Figure 1: Chromosomal Map of the region of yeast chromosome 7 containing the genes PCP1 and YGR102C. (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/ORFMAP/ORFmap?sgdid=S0003334>).
This gene is located on yeast chromosome VII. The systematic name for PCP1 is YGR101W, and other aliases are MDM37 and RBD1. (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=PCP1>).
Click here for the genomic DNA sequence. It consists of 1041 base pairs.
Click here for the Entrez-Protein link for PCP1. It is 347 residues long.
More sequence and translation information for PCP1 is available here.
A PDB search produced no results.
The gene product of PCP1 is a protein called rhomboid protease.
The Biological Process, Biological Function and Cellular Component as defined by Gene Ontology are as follows. (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=PCP1>).
Molecular function: peptidase activity i.e. catalysis of the hydrolysis of peptide bonds.
Biological process:
mitochondrion organization and biogenesis i.e. assembly and maintenance of a mitochondrion.
Cellular component: mitochondrion (in mitochondrial inner membrane).
According to the SGD, the phenotype is as follows. (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/phenotype/phenotype.pl?feat=PCP1&type=locus>).
Exhibits growth defect on a non-fermentable (respiratory) carbon source.
Null: lack of Ccp1 maturation, slow growth on non-fermentable media.
The hypothetical ORF YGR102C is located on chromosome VII in close proximity with PCP1. Its cellular component, biological process, and molecular function are unknown. (SGD, 2003; <http://db.yeastgenome.org/cgi-bin/SGD/locus.pl?locus=YGR102C>).
Click here for the genomic DNA sequence. It consists of 552 base pairs.
Click here for the Entrez-Protein link for YGR102C. It is 183 residues long.
Predictions on the nature of the protein can be made using Kyte-Doolittle hydropathy plot.
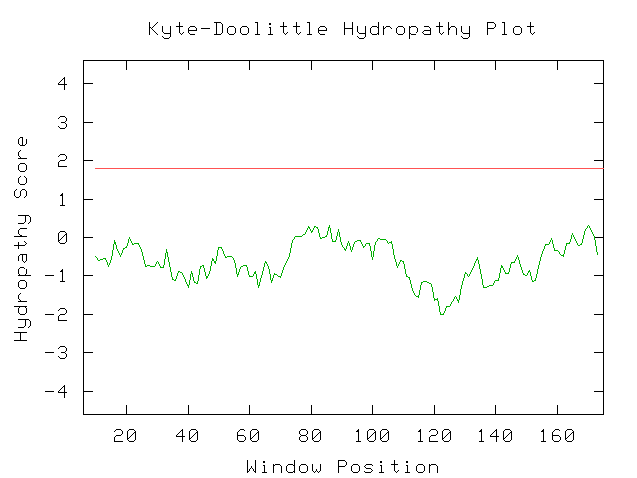
Figure 2: Kyte-Doolittle Plot
When the window size is 19, no parts of the protein have a hydropathy score greater than 1.8. Thus we can deduce that the protein does not have any trans-membrane regions but may be located somewhere in the cytoplasm.
We can also compare the genomic DNA and protein sequences to known sequences of other genes.
A BLASTn search yielded the following:

Figure 3: Results of BLASTn search.

This indicates that the gene showed significant similarity to two other genes.
YGR103W in Saccharomyces cerevisiae.
U67492 in Methanocaldococcus jannaschii.
BLASTp also produced a good match.
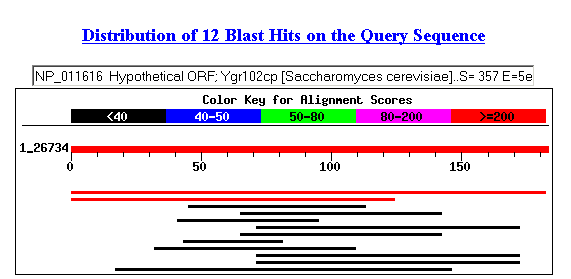
Figure 4: Results of BLASTp search.

The potential matching protein was an unnamed yeast protein product (accession number: CAA97107). It is encoded by the gene YGR103W, the same one that BLASTn yielded.
Click here to learn more about YGR103W at the SGD. Also called NOP7, it is a nucleolar protein present in purified ribosome assembly intermediates. It is required for rRNA processing; required for essential steps leading to synthesis of 60S ribosomal subunits. However, its molecular function is unknown. Notice in Figure 1 that NOP7 is located in close proximity to YGR102C on the same chromosome.
Since YGR102C has a DNA sequence similar to that of NOP7, and it also encodes a protein that is very similar to the one encoded by NOP7, we can hypothesize that YGR102C is a nucleolar protein with a potentially similar function.
Send questions, comments and suggestions to: pakarnik@davidson.edu