- This web page was produced as an assignment for an undergraduate course at Davidson College -
My Favorite Yeast Genes
EST1 and YLR243W
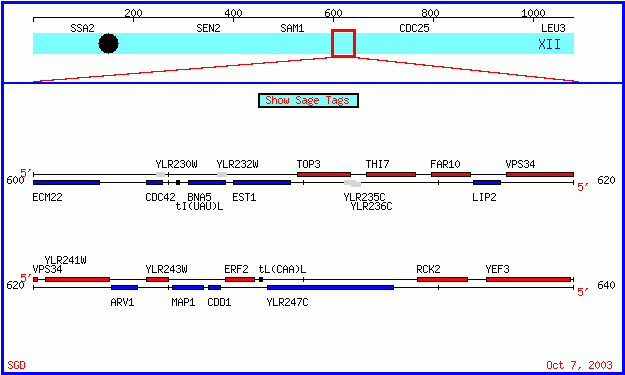
EST1 and YLR243W are genes found in the unicellular fungi Saccharomyces cerevisiae. As of October 2003, YLR243W is unannotated - that is, specific information regarding the gene's molecular function, biological process, and cellular localization is largely unknown. EST1, also known by the systematic name YLR233C, has been well characterized. EST1 and YLR243W are located in close proximity to each other on S. cerevisiae chromosome XII. The map below, produced by the Stanford Saccharomyces Genome Database indicates the spatial relationship of these two genes. Note that YLR243W is located on the Watson (red) strand and EST1 is located on the Crick (Blue) strand. In this webpage, we will investigate these genes using available online databases.
EST1
Oddly enough, NCBI does not list an updated sequence for the wild-type Est1 gene (also called YLR233C or L8083.15 by Swiss-Prot). The sequence is contained within the larger complete chromosome XII sequence (NC_001144). Particular sequence and translation information for the gene can be found at the Stanford Saccharomyces Genome Database. Click on the Yellow SGD bubble above to visit the homepage for the Est1 locus. A mutant form of the gene is listed in NCBI (J04849). By following the PubMed link to the paper related to this mutation (Lundblad and Szostak, 1989), we read about the first characterization of the locus. It was found that the telomeres in mutant yeast are shortened and the yeast are more susceptible to chromosome loss. For the observed mutant phenotype, the gene was named “Ever Shorter Telomeres” and abbreviated Est1. Subsequent papers identified two additional EST genes which are involved in telomerase activity (Linger et al., 1997). More recently, researchers have implicated Est1 directly in telomere-binding during late S phase which influences cell cycle regulation (Taggart et al., 2002).
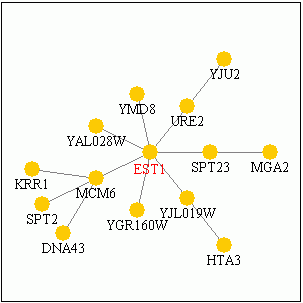
It is generally understood that the biological process of EST1 is to associate with telomerase and assist in telomere maintenance. For a background on telomeres and telomerase, visit Kimball's Biology Pages. Telomeres are part of chromosomes which are located in the nucleus, and the EST1 protein is also found in that cellular component. The complex formed with telomerase is a ribonucleoprotein enzyme complex called "telomerase holoenzyme complex" which associates with many of the proteins in the picture below. On the molecular level, it is generally understood that EST1 has an RNA binding domain to bind with the telomerase ribonucleoprotein and a DNA binding domain to bind with the chromosomal telomeres. More specific interactions, however, have met with some dispute in the scientific community.
Today, researchers remain unclear as to the nature of EST1 protein and the specific events regulating telomere length. Findings have allowed researchers to devise three separate models to describe Est1 (Taggart and Zakian, 2003). Why is EST1 function so difficult to describe? After all, the gene was identified over 14 years ago; surely we have had ample time to elucidate the molecular connections involved. In fact, it has been done – and the data give good reason why the cellular mechanisms are not perfectly understood. Curagen Corporation makes proteomics technology and from their website we can generate a simplified network of molecules related to EST1. Click on the diagram below, and then explore the associated proteins by clicking on them within the PathCalling website. Notice how many iterations exist just within 2-3 nodes of the source. This level of complexity is a good reason why more work is needed to fully understand the intricate functions of EST1.
YLR243W
YLR243W
is an unannotated yeast gene located on chromosome XII. If you click the yellow SGD bubble above, you
can see that the molecular function, biological process, and cellular component
of the expressed gene are all listed as unknown. In fact, the only description of YLR243W
given is, “Protein required for cell viability.” This statement is based off of the (nearly)
genome-wide ORF deletion study conducted at the
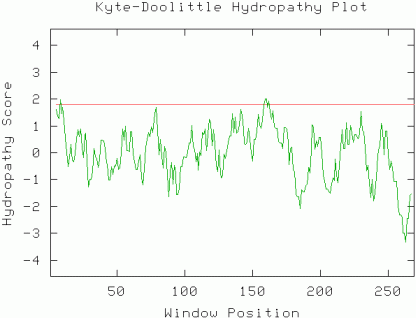
The gene sequence can be viewed at NCBI as part of the complete chomosome XII sequence and the protein conceptual translation can be viewed at NP_013344.1. Note that this is not an experimentally determined protein sequence, but a predicted translation based on the gene ORF. The ORF is 819 base pairs and the predicted protein is 272 residues long. Initial predictions on the nature of the protein may be deduced by a Kyte-Doolittle hydropathy plot. Almost the entire protein has a hydropathy score >1.8 (window=9). This suggests that the protein does not have an integral membrane domain and may be located in the cytoplasm.
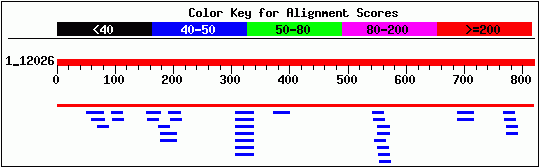
The next step in characterizing this gene is to compare the DNA and protein sequences to known sequences of other genes. A BLASTn search did not yield any significant similarity with other known genes. In the alignment below, the top red line is the input YLR243W sequence, the second red line indicates the match with the Saccharomyces chromosome XII contig, and the blue lines below represent known sequences of slight homology. BLASTn did not return any homologous sequences with E-value <1.5.
BLASTp, however, did yield many potential hits; 42 proteins had E-value >.001. These potential matching proteins were from a wide range of organisms (from Arabidopsis to Drosophila to humans), yet all contained a similar description. Namely, they all mentioned that the protein was a “conserved hypothetical ATP binding protein.” Again, the specific function is only hypothetical, but the fact that the protein is found in many organisms does bespeak its importance in biological processes. The Swiss-Prot database also mentions that the molecular function is ATP binding, but that is inferred from electronic annotation (Q06543). The most closely aligned human protein, x0004 hypothetical ATP binding protein, can be found at NP_057385. The Yeast genome database lists mammalian orthologs and the closest listing in this database is a Human ATP/GTP binding protein (AAC50780).

A conserved domain search validated the potential for ATP-binding function.

Since no structure exists in NCBI Structure or PDB, predictions can be made regarding gross protein structure by Predator or Invitrogen. Both predicted structures contain large sections of random coil or alpha helix.
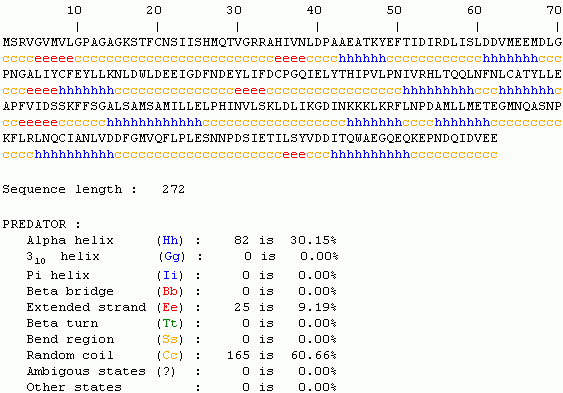

We have determined that the yeast unannotated proteinYLR243W is essential for yeast survival, has conserved orthologs in many species, is probably cytosolic, appears to be comprised of many coils and helixes, and has a predicted ATP-binding domain.
References
Giaever G, Chu AM, Ni L, Connelly C, Riles L, Veronneau S, Dow S, Lucau-Danila A, Anderson K, Andre B, Arkin AP, Astromoff A, El-Bakkoury M, Bangham R, Benito R, Brachat S, Campanaro S, Curtiss M, Davis K, Deutschbauer A, Entian KD, Flaherty P, Foury F, Garfinkel DJ, Gerstein M, Gotte D, Guldener U, Hegemann JH, Hempel S, Herman Z, Jaramillo DF, Kelly DE, Kelly SL, Kotter P, LaBonte D, Lamb DC, Lan N, Liang H, Liao H, Liu L, Luo C, Lussier M, Mao R, Menard P, Ooi SL, Revuelta JL, Roberts CJ, Rose M, Ross-Macdonald P, Scherens B, Schimmack G, Shafer B, Shoemaker DD, Sookhai-Mahadeo S, Storms RK, Strathern JN, Valle G, Voet M, Volckaert G, Wang CY, Ward TR, Wilhelmy J, Winzeler EA, Yang Y, Yen G, Youngman E, Yu K, Bussey H, Boeke JD, Snyder M, Philippsen P, Davis RW, and Johnston M. (2002). Functional profiling of the Saccharomyces cerevisiae genome. Nature. 418: 387-91. (PubMed).
Linger J, Cech TR, Hughs TR, and Lundblad V. (1997). Three Ever Shorter Telomerase (EST) genes are dispensable for in vitro yeast telomerase activity. PNAS. 94: 11190-5. (PubMed).
Lundblad V and Szostak JW. (1989). A mutant with a defect in telomere elongation leads to senescence in yeast. Cell. 57: 633-43. (PubMed).
This page created by: Gray Lyons ![]()
Return to My Genomics Homepage
Return to the Genomics Course Homepage
Return to the Davidson College Homepage