this site is maintained by Nona Poulton under the supervision of Dr. A. Malcolm Campbell as part of the course requirements of Genomics at Davidson College, Davidson NC
Characterization of RPB8 and YOR223W Protein Interactions
GO Information: What is known so far about the RPB8 protein
(see web assignment #2)
Molecular Function: RPB8 encodes part of DNA-dependent RNA
polymerase, where it functions as one subunit of RNA polymerase I, II, and III.
It aids in the catalysis of DNA-template-directed extension of the 3'- end of
an RNA strand, one nucleotide at a time.
Cellular Component: RNA-polymerase initiates RNA synthesis
from DNA inside the nucleolus.
Biological Process: Synthesis of RNA from DNA template via
RNA-polymerase (I, II, and III), of which RPB8 encodes a subunit. Synthesis
starts at the RNA-polymerase-specific promoter.

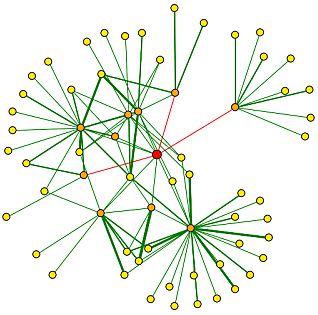
Figure 1: This image shows RPB8s ineractions in a network form. The DIP website compiles a bunch of information. The legend can be seen below. RPB8 is the main dot in the centre (red). It has many connections, some are red, which means they are unverified connections. By clicking on the yellow dots around RPB8, I found (not surprisingly) that the proteins that interact with RPB8 are all RNA polymerase subunits.

Figure 2: MIPS results for RPB8. MIPS provides information about yeast protein function, including the above table. It shows that RPB8 (YOR224c) interacts with many proteins. When I clicked on them, they were all other RNA polymerase subunits, consistent with the above data from DIP.
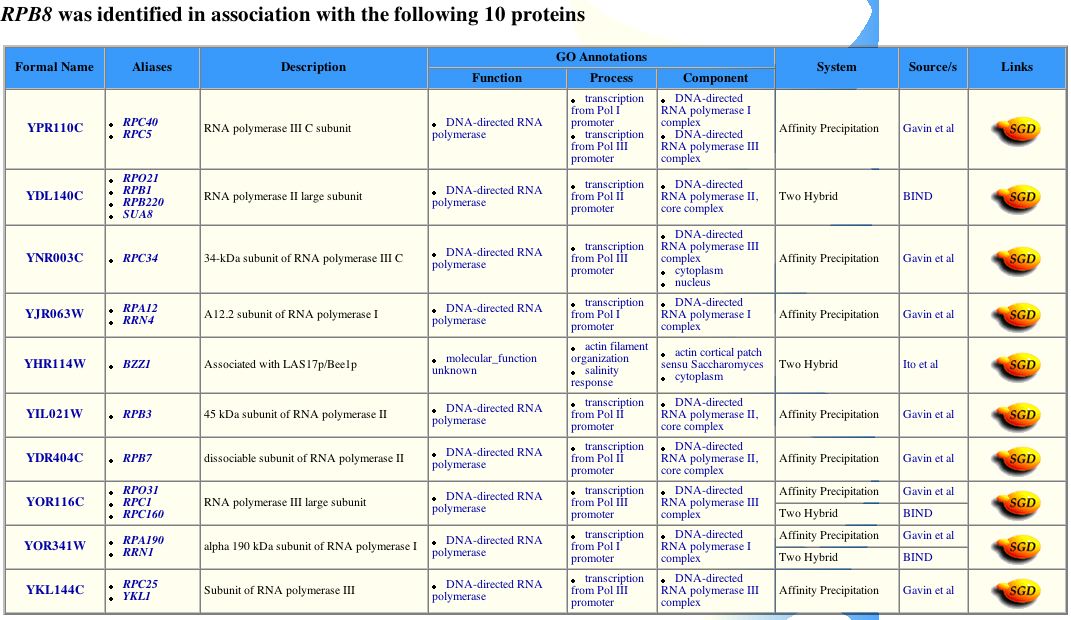
Figure 3: The Yeast Grid provides information on genetic and physical interactions of yeast proteins developed in Tyers Group at the Samuel Lunenfeld Research Institute at Mount Sinai Hospital.The figure shows that there are 10 protein interactions, all RNA polymerase subunits (consistent with all the above information). The YHR114W protein (not listed as an RNA polymerase subunit) seems out of place. When I followed the link, it turns out that the protein interacts with many proteins (197), none of which are similar. It is an actin patch, which explains its interactions. It does not make sense, however, for this protein to interact with RPB8. Further studies should be done to find out what this interaction means or whether it is a mistake in the database.

Figure 4: Pathcalling results show no interactions (see lonely RPB8 above).
![]()
Figure 5: PROWL is a database that allows the user to calculate the molecular weight and isoelectric point of a given protein. The results for RPB8 are above. The website also provides links to other databases (most discussed below), but these links provided no information for RPB8.
Yeast Protein Linkage Map Data: This is another yeast 2-hybrid system with a list of prey proteins and their predators. A search for RPB8 retrieved no results, however.
TRIPLES: (TRansposon-Insertion Phenotypes, Localization, and Expression) website provides information on protein interactions as well. In this case, researchers at Yale produced transposable elements, which were used to disrupt genes, producing certain phenotypes under various growth environments. Disruptions made in RPB8 did not provide any information about the function because the tested did not differ from wildtype. RPB8 probably does not respond to any of the experimental conditions they tested.
Swiss 2-D Plot: This database provides 2-d electrophoresis plots of various proteins, which would be useful information if RPB8 was present.
Yeast Resource Centre's Yeast Two-Hybrid Analysis: This website contains information about protein to protein interactions in S. cerevisiae. The data was obtained by yeast two-hybrid analysis (information on website). It does not contain information on RPB8, however. The supplemental data (published since the page waas created) also did not contain any information on RPB8.
PDB results show no new information for RPB8 and its interactions.
Benno
Figure: This .pdf file is a lage comprehensive protein interaction map.
It shows many interactions, but a search for RPB8 returned no results.
Conclusions: The main conclusions to be made about RPB8 protein interactions are that the protein interacts with other RNA polymerase subunits, which is not surprising. In fact, these interactions could be predicted not using any of the above methods. In light of these findings, there is no reason to reject the given DO information provided above.
GO Information: What is known so far about the YOR223W protein
(see web assignment #2).
Molecular Function: We know that it is not only crucial for
cell viability but that it is probably required for cellular organization processes,
especially those that are important to the life of the cell.
Cellular Component: YOR223W is an integral membrane protein
and is probably involved in cell membrane organization and biogenesis.
Biological Process: The YOR223W gene product is involved in
cellular organization.
DIP: This website provided no information for YOR223W. This could be because it is unannotated with no known function.
MIPS: Provided no results for YOR223W.
Yeast Grid: Search for YOR223W returned no interactions.
TRIPLES: Produced no meaningful results. YOR223W was disrupted in two places, both within the ORF. The conditions had no effect and the results looked like wild type in all cases.
Swiss 2-D Plot: No information for YOR223W.
Yeast Resource Centre's Yeast Two-Hybrid Analysis: Does not include YOR223W in prey or predators.
PDB results show no new information for YOR223W and its interactions.
Benno Figure: The find function of adobe did not retrieve any hits for YOR223W, so I assume it is not included in this map.
Conclusions: Proteomics has a long way to go. The YOR223W (along
with RPB8) does not show up in any of the databases. This could be because not
all proteins have been used as predators or that the technology is so new that
there has not been time yet to analyze all the data. This protein is also very
much unannotated. There is hardly anything known about it, which leads me to
believe that it is not involved in any of the traditional experimental pathways
(glycoloysis, for example), or is simply one of the many proteins that no one
knows anything about. After finding no new results using the databases discussed
above, I am not able to add anything new to my GO predictions. The experiments
below may help to uncover some things about YOR223W, however.
Based on previous information, YOR223W is an integral membrane protein. To test this further and perhaps get more meaningful results (other than the hydropathy plot guesses) pronase, a protein digestion enzyme, can be used. Because pronase digests proteins, it is possible to compare the presence of YOR223W with or without pronase treatment. If YOR223W is not present after pronase treatment and is present when the cell is intact, then we can reasonably conclude that YOR223w is most likely a membrane protein. This limits its functions as well, which could provide further insights to possible experiments.
It may be useful to use yeast two-hybrid experiments to test my prediction made above. It is thought that YOR223W may be involved in cell membrane organization and biogenesis, so there could be two experimental tests there. Known proteins involved in membrane organization could be used as prey to get meaningful results. Interactions (or lack of interactions) would provide information on the protein's function.
To test whether YOR223W undergoes phosphoylation, it may be useful to use the polymerized chip PDMS, as Mike Snyder has done. There are 122 known kinases in yeast, which could be incubated with YOR223W to see interaction. If radioactive ATP was used as a substrate, radioactive P would be taken up by YOR223W and would be bound to the chip, which could then be assessed for radioactivity. If the protein is phosphorylated by one of the kinases, then radioactivity would be detected on the chip after all substrates had been washed off.
References:
Yeast Protein Linkage Map Data
Yeast Resource Centre's Yeast Two-Hybrid Analysis
© Copyright 2003 Davidson College, Davidson, NC 28035