This webpage was produced as an assignment in an undergraduate course at Davidson College
My Favorite Yeast Protein
Annotated: ERG25
From my previous websites on ERG25 I noted that this gene and its protein product are involved in the ergosterol biosynthesis pathway. C-4 methyl sterol oxidase, as the protein is called, is involved in the process of creating sterols from simpler compounds. I continued my ERG25 search by looking at protein function. I began by looking at Swissprot to determine if there was any more information on ERG25. The image below confirms my earlier finding of the gene's function but also notes that it requires iron as a cofactor in its synthesis of sterols. It also supports my earlier finding that the protein is a membrane protein localized in the endoplasmic reticulum. Swissprot also notes that the active site might be located in the histidine box.

Figure 1 - Information about ERG25 from Swissprot
I then searched MIPS for more information about the protein produced by ERG25 (Figure 2). MIPS notes that ERG25 has similarity to a C. elegans hypothetical protein F49E12.9. This similarity might help in the indentification of the C.elegans protein, but that's a side note. The page also notes the protein's isoelectric point of 8.17 and its molecular weight of 36.4793 kDa. This site also confirms the proteins role in metabolism, specifically lipid metabolism. MIPS also notes that deletions in this gene is lethal lending to the conclusion that the step ERG25 performs in removing two C-4 methyl groups leading to ergosterol is essential to cellular functioning.
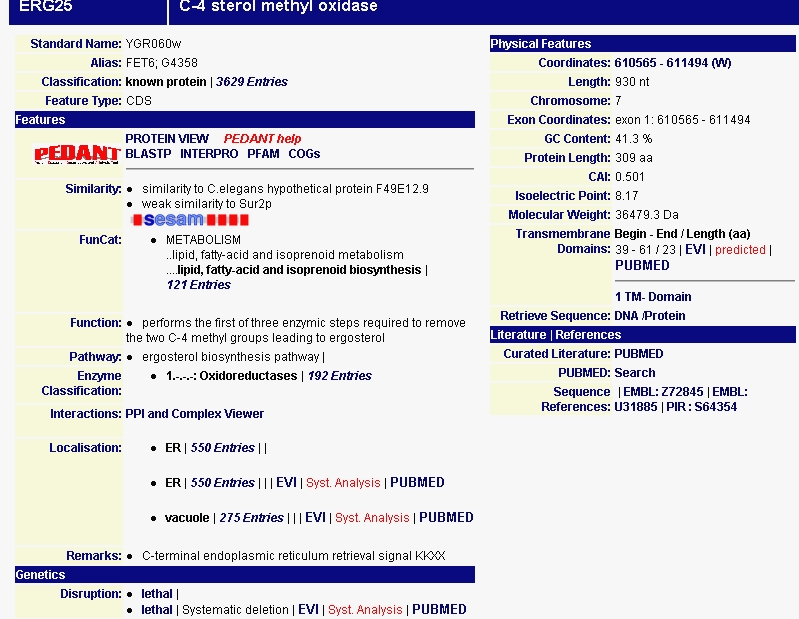
Figure 2 - Data on ERG25 from the MIPS database
When I searched the Yale Gerstein lab's webpage I found some confounding data that I'm unsure how to interpret (figure 3). They predict ERG25 to be located in the cytoplasm. Every other source specifically states the protein is located in the membrane of the endoplasmic reticulum. There is a chance that their finding are old and that the closest they could predict the protein's localization was somewhere in the cytoplasm. I found the inconsistency of the data available interesting and worth noting.
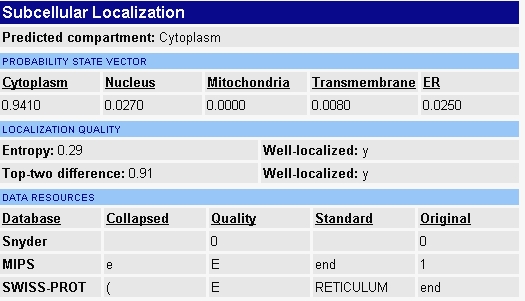
Figure 3 - Predicted location of ERG25 from Yale Gerstein Lab.
The diagram below from the DIP database shows ERG25 as the red large dot. Two of the three proteins located above ERG25 are non-annotated membrane proteins. These proteins are probably also involved in ergosterol or steroid metabolism. The protein below ERG25 is called Securin and, strangely, is a regulatory protein involved in chromosome stability. I say strangely because this appears to have little connection to ERG25's function in ergosterol biosynthesis. Swissprot notes that this protein acts in the nucleus during mitosis to block separase; it ultimately helps the segregation of the chromosomes. All of the proteins below securin are related to this process during cell division. This is confusing. Why would these proteins displayed as integral to ERG25's function when they act on completely different cellular processes?
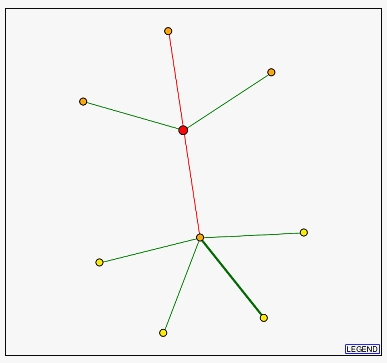
Figure 4 - Diagram of proteins related to to ERG25 from DIP
I went to MIPS protein-protein interaction page to discover whether these other proteins interacted with ERG25 (figure 5). This search showed that these proteins do not interact with ERG25. ERG25 interacts with three proteins ERG26 or C-3 sterol dehydrogenase, ERG27 or 3-keto sterol reductase, and ERG28 another protein involved in ergosterol synthesis. As their names imply these three proteins are also involved in the ergosterol biosynthesis pathway. MIPS notes that ERG25 works in concert with ERG26 and ERG27 to catalize the removal of two methyl groups at the sterol C-4 position. MIPS showed no protein complex interactions from their yeast two-hybrid studies. Unfortuantely, my search of the Benno PDF files were all fruitless. This protein is fairly well understood. It's location and function are known, and researchers have also studied ERG25's interaction with other proteins in the ergosterol biosynthesis pathway. Even when other steps of the ergosterol pathway are unclear ERG25, ERG26, ERG27 are well understood in their collective function of removing two methyl groups at the sterol C-4 position.
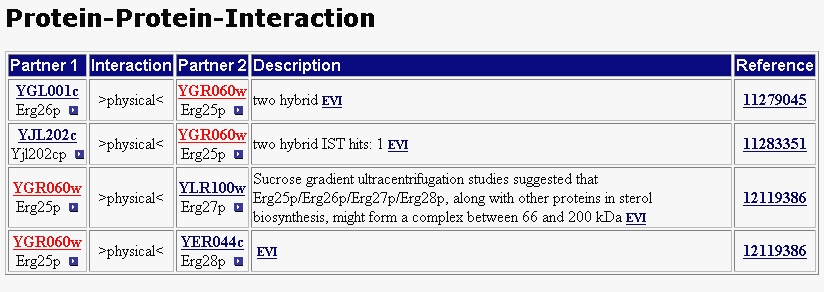

Figure 5 - Protein-protein interaction and protein complex from MIPS.
MIPS - http://mips.gsf.de/genre/proj/yeast/index.jsp
DIP - http://dip.doe-mbi.ucla.edu/dip/Search.cgi?SM=3
Yale Gerstein Lab - http://bioinfo.mbb.yale.edu/
The Genomics Place - Benno PDF Files - http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/chapter6/deluxe.html
YRC Two Hybrid Analysis - http://depts.washington.edu/%7Eyeastrc/th_11.htm
Contact: Alex Trzebucki