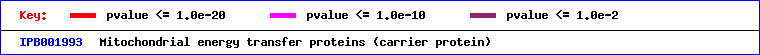
In the figures above, using the eMOTIF database, one can see that the structure of the protein is similar to many other mitochondrial transfer proteins.
My Favorite Yeast Genes
This webpage was produced as an assignment for an undergraduate course at Davidson College
Introduction:
The purpose of this web page is to examine the functions of two genes from the yeast species Saccharomyces cerevisiae. The first gene is annotated, which means information is already known about the molecular function, biological process, and cellular components. The second gene however, is non-annotated: using several different resources, I will attempt to predict as many characteristics of this unkown gene as possible.
Annotated gene: MTM1
Information on this gene can be found at the Saccharomyces Genome Database (SGD)
Information about the protein from this gene can be obtained at NCBI, with accession number gi:6321696
For the complete nucleotide sequence, accession number gi:50593213 can be used.
Molecular Function:
As stated in the orginial paper characterizing this gene (Luk, et al. 2003), MTM1 stands for: manganese trafficking factor for mitochondrial SOD2. The protein is involved in a metallochaperone activity, assisting in the delivery of manganese ions to the mitochondria.

In the figures above, using the eMOTIF database, one can see that the structure of the protein is similar to many other mitochondrial transfer proteins.

The above figure was obtained using the NCBI BLASTp database. This image again confirms the domains present in the MTM1 protein.
Biological Process
The MTM1 gene encodes a protein that represents a member of the mitochondrial carrier family (MCF), however this particular protein is not a general manganese transporter, but is specific to SOD2, superoxide dismutase.
Cellular Location:
MTM1 is a mitochondrial protein and is found in the mitochondrial inner membrane. This location places the Mtm1p (protein formed form the MTM1 gene) very close to the site of SOD2 translocation into the matrix. This allows for the possibility that the manganese insertion by the Mtm1p into the SOD2 receptor takes place as the SOD2 protein enters the mitochondrial matrix (Luk, et al. 2003). The protein also has 4 transmembrane domains as predicted by the Saccharomyces Genome Database, this information can be found at Predicted Features.
Chromosomal Location:
MTM1 is located on the Crick strand of Chromosome 7 in Saccharomyces cerevisiae.
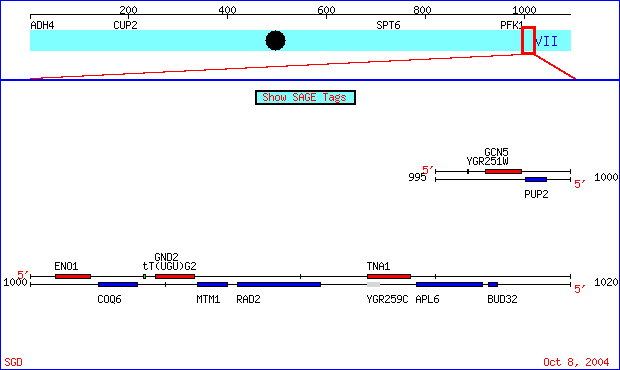
Above figure indicates location of MTM1 gene in relation to surrounding genes on Chromosome 7. Non-annotated gene YGR259C can be found slightly to the right of the MTM1 box. The blue box associated with MTM1 indicates that the gene is a Crick strand ORF. The gray box for the non-annotated gene indicates a dubious ORF with no known functions.
Complete nucleotide coding sequence (does not include untranslated sequence regions or bases, such as introns):
ATGAGTGATCGCAATACAAGTAACAGCCTGACCTTAAAAGAACGAATGTTAAGTGCCGGT GCTGGATCAGTACTCACTTCTTTGATTTTGACACCGATGGATGTAGTGAGAATACGGCTG CAACAGCAGCAGATGATTCCTGACTGTTCATGTGATGGGGCTGCCGAAGTGCCCAACGCC GTTAGCTCAGGCTCCAAAATGAAGACATTCACCAACGTCGGAGGACAGAACTTGAACAAC GCAAAAATATTTTGGGAGAGTGCTTGTTTTCAAGAACTGCACTGTAAAAATTCATCGTTG AAGTTCAACGGTACTTTAGAAGCATTCACGAAAATTGCCAGCGTCGAAGGTATTACAAGT TTGTGGAGGGGTATTTCTTTGACCCTATTAATGGCTATTCCTGCCAACATGGTTTATTTT AGTGGTTATGAATATATAAGAGACGTTTCTCCTATAGCCTCGACGTACCCAACATTAAAC CCCCTCTTTTGTGGTGCAATTGCGAGGGTATTTGCAGCAACAAGTATTGCACCTTTAGAA TTGGTTAAAACTAAACTGCAAAGTATACCGAGATCATCAAAGTCCACAAAGACATGGATG ATGGTTAAAGATTTATTAAACGAAACGAGGCAGGAAATGAAAATGGTGGGTCCTTCTCGG GCCCTCTTCAAAGGTTTGGAGATCACTCTGTGGAGAGACGTTCCGTTTAGTGCAATATAT TGGAGTTCCTATGAACTTTGCAAGGAAAGACTATGGCTAGATTCTACTCGATTTGCATCT AAGGACGCAAACTGGGTCCATTTTATAAACAGTTTTGCCAGTGGTTGCATAAGTGGGATG ATAGCTGCTATATGCACACACCCTTTTGATGTCGGTAAGACGAGATGGCAAATATCCATG ATGAACAATAGCGATCCGAAAGGCGGCAATAGGTCTAGAAACATGTTCAAGTTCTTAGAG ACTATATGGCGTACAGAAGGTCTTGCGGCCCTCTACACGGGCCTGGCAGCCAGAGTAATT AAGATACGCCCAAGTTGCGCCATCATGATATCTAGTTATGAGATCTCCAAAAAAGTATTT
GGAAACAAATTGCATCAGTGA

This figure shows the number of homologs in other organisms that have sequence alignment with the MTM1 gene. As you can see from the image below, there is a human homolog to this gene as well.
In the figure above, one can see that the Homo sapiens and Mus musculus genes are closely related to the Saccharomyces cerevisiae. To view alignments with homologs from other organisms, please see this site. This image was generated using the PSI-Blast analysis of MTM1 on the SGD.
The MTM1 gene is also associated with several mutations: when the human homolog is mutated, the result is a disease known as myotubular myopathy. For yeast, the mutation leads to severe growth defects. This is due to the essential interaction between MTM1 and SOD2: "SOD2 plays a critical role in guarding against mitochondrial oxidative stress and is essential for survival of many organisms" (Luk, et al., 2003). When Mtm1p is mutated, manganese cannot be delivered to SOD2 and the cell accumulates toxic levels of manganese.
Non-annotated gene: YGR259c
This is a hypothetical Open Reading Frame (ORF) very close to the previously explained gene MTM1 on Chromosome 7. However, non-annotated means that the molecular function, biological process and cellular components are not yet known. Therefore, it is necessary to use research techniques in order to predict the function of this ORF.
There is currently no information about the gene in the public domain: when searching the NCBI Gene Database, no matches were found.
Nucleotide Sequence:
ATGGGGTCGAAAAGAACATACGTCGCGAAAAAAACTGTGACACATGTGTTATATTGCGTT CCTACAAGGTGGATATCCTTGGATAGACCTGCAACTTCTGCGTTACCAATATTCGATTTA TCCAGGTTAGAAAGGAAATATAGCATCCCCATAAGAGGAATCAAGTATATATCCATTTTG CGCAAGACACGCTTTTCCATTGGATGGGAAGGGGAGGGGATGCCGTCATCATCCGTGGTA GTAGCCATAATTTCCCGTTCGGTGGCAAGTTCCGACTTCTTGTCATGGCTGCGATTATGC AAGCTAGCGTCATGGCCTTCATCTTCCTGGAACGTGACTTCCGTAGGCTTTTCTTCTGAT CCATCATTGGTGGGTGATATGAAAAGCACATCATCCACCAAATGCTTAGGTGACTCCATT GTAAATTTGTTGCTCATCTAG
Because the function of this ORF is unknown, it is not possible to tell whether all of this is a coding sequence, or if there are untranslated regions within the sequence. This sequence is shown in the 5'-3' direction of the Crick strand.
The entry into the NCBI sequence database can be found with accession number gi:1323471
Protein information
Amino Acid Sequence
1 mgskrtyvak ktvthvlycv ptrwisldrp atsalpifdl srlerkysip irgikyisil
61 rktrfsigwe gegmpsssvv vaiisrsvas sdflswlrlc klaswpsssw nvtsvgfssd
121 pslvgdmkst sstkclgdsi vnlll
According to the Yeast Genome Database, the protein resulting from this open reading frame would have one transmembrane domain. The protein is only 146 amino acids long, and the accession number is CAA69081.1
This transmembrane domain can then be confirmed with a Kyte Doolittle Hydropathy Plot:


When a PSI BLAST was done using the YGR259c sequence, no homologs were found in any other species. This could be due to the fact that the function is unknown, and because no putative conserved domains were discovered, it would be difficult to match the predicted function with any other species.

When a BLASTp was done using the NCBI website, no significant matches were found with other protein sequences in the database. An E value of 4.2 indicates that the match was merely coincidence. This is important because there are only 146 amino acids in the protein, but they are in such an order that no other species has a protein quite like it. Thus this could be a gene with a function related solely to the necessities of yeast life.

Using PREDATOR to predict the Secondary structure of the protein, we see that the protein is almost completely of random coils, however there are some areas of alpha helix and extended strands.
Prediction:
Due to the fact that so little is known about this gene, it is necessary to use the information gathered to make a prediction. Since the protein only significantly matched with itself, this means there are no similar proteins in other organisms. While the location of the ORF is known (see Chromosome location picture in MTM1 section), the localization of the protein within the yeast is unknown. There is a transmembrane domain, thus the protein can be found in a membrane of the cell. The Predator information tells us that the protein has many random coils: however just because they are named 'random coils' does not mean they are random. The protein could be folded in such a way as to create a ligand binding location for another yeast protein. Thus I believe the protein is a receptor in a membrane for a pathway unique to yeast, for example the yeast budding pathway.
References:
Due to the recent discovery of the MTM1 gene, there is only one paper written specifically about this gene. The non-annotated gene has no references because no functions have been discovered for the ORF.
Luk, E., Carroll, M., Baker, M., Cizewski Culotta, V. 2003. "Manganese activation of superoxide dismutase 2 in Saccharomyces cerevisiae requires MTM1, a member of the mitochondrial carrier family". Proc. Natl. Acad. Sci. USA; 100, 10353-10357.
Many Databases were used in researching this website:
Megan McDonald's Genomics Homepage
Send questions or comments to: Megan McDonald