This web page was produced as an assignment for an undergraduate course at Davidson College
Interpreting the Expression of Yeast Genes CYT1 and YOR066W using Microarrays
CYT1
In a previous web page, the gene CYT1, an electron transporter in the mitochondrial inner membrane in Saccharomyces cerevisiae, was examined. To better understand the function of this gene, specifically its expression pattern under varrying conditions and which other genes may have similar functions, data from the Expression Connection website was analyzed. The spectrum below shows the scale with which microarray clustered data was analyzed.
* next to GO terms indicates that only one of the GO annotations for that GO aspect is listed on this page, for this ORF (Balakrishnan, 2004).
Scale: (fold repression : fold induction)

There is a lot of available information in the form of clustered data. In order to interpret CYT1 expression more objectively, some of the clustered results were computed into base 2 logarithm graphs. I chose to interpret the yeast expression studies for which base 2 logarithmic graphs had been generated. Each experimental set of data was clustered according to genes that had a Pearson correlation coefficient > 0.8. Values close to -1 exhibit strong but negative correlation, values close to 0 are interpreted as having no significant correlation and values between 0.8 are used here and exhibit strong, positive correlation between genes. The expression patterns of both unannotated and functionally established genes were correlated to CYT1.
Expression Pattern 1: during sporulation
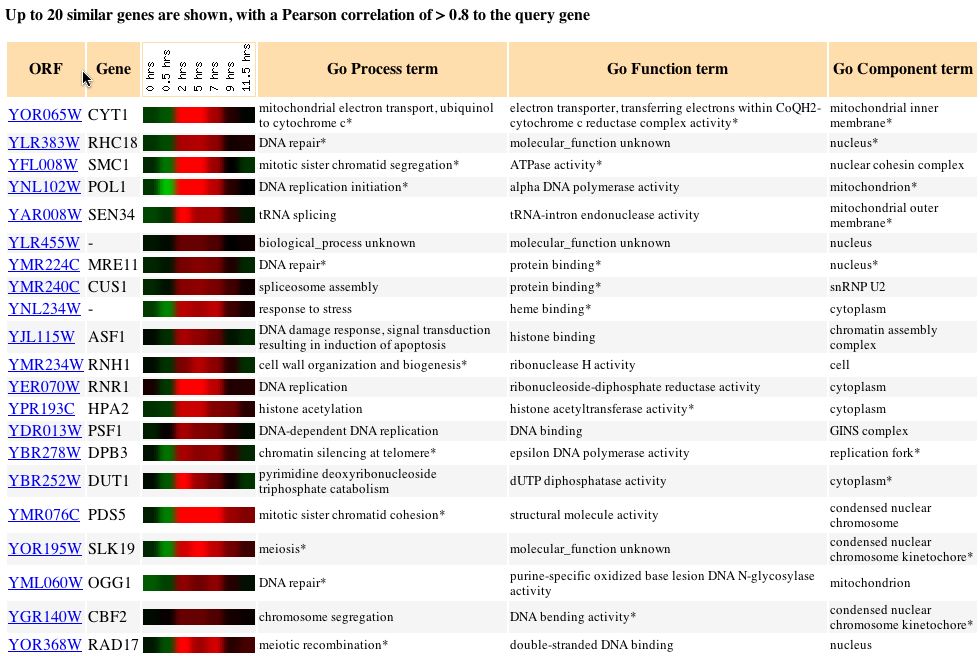
Fig. 1. Cluster pattern for sporulation over time (SGD, 2004).
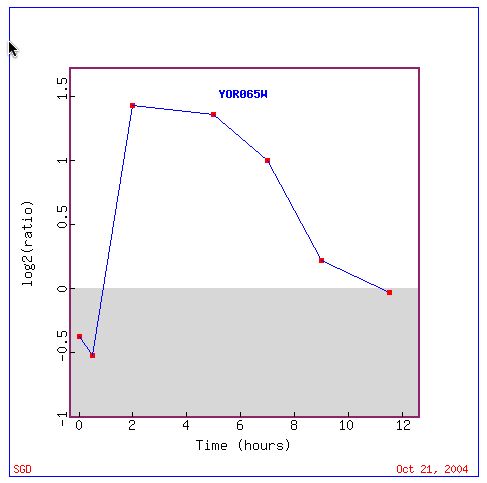
Fig. 2. Strong induction is seen in these genes between two and seven hours. Note that genes are initially repressed (SGD, 2004).
In the expression pattern of sporulation, asexual reproduction by budding yeast, induced genes share involvement in mitotic activity. The period of time for which induction of occurs (Fig. 2) suggests that there is a window within the cyclical sporulation process during which the clustered genes are most actively transcribing proteins, such as CYT1.
Expression Pattern 2: the diauxic shift
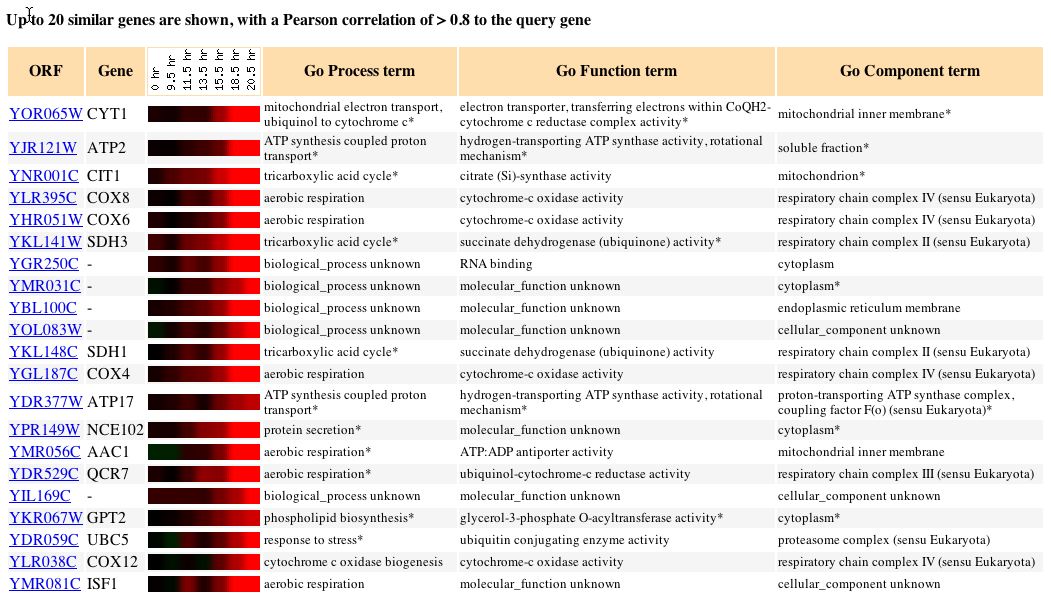
Fig. 3. Clustered data for CYT1 during diuaxic shift over time (SGD, 2004).
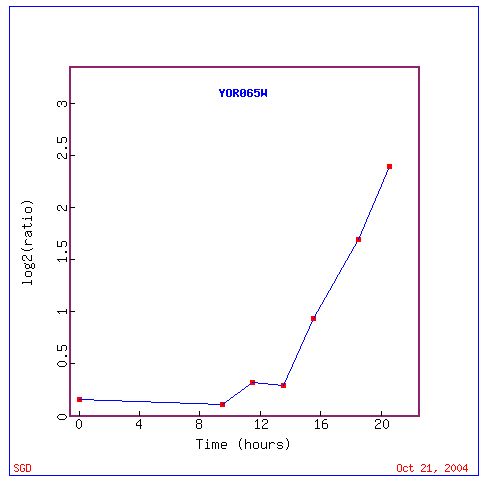
Fig. 4. Expression of CYT1 is increasingly induced from 14 - 20 hours (SGD, 2004).
The diauxic shift occurs in yeast when anaerobic fermentation of glucose, the preferred method of yeast metabolism provided there is enough glucose, shifts to aerobic respiration of ethanol, which is present in high concentrations once the carbon source in glucose has been exhausted (DeRisi, 1997). Since the whole metabolic pathway changes, the pattern of gene expression is expected to change. The data show that the pattern of CYT1 expression does change, CYT1 becomes more induced over time. The data suggests that the CYT1 mitochondrial electron transport protein is involved in the metabolic pathways associated with diauxic shift in yeast.
The following two experiments look at gene expression in response to alpha-factors. Alpha-factors are a signaling molecule involved in yeast sexual reproduction.
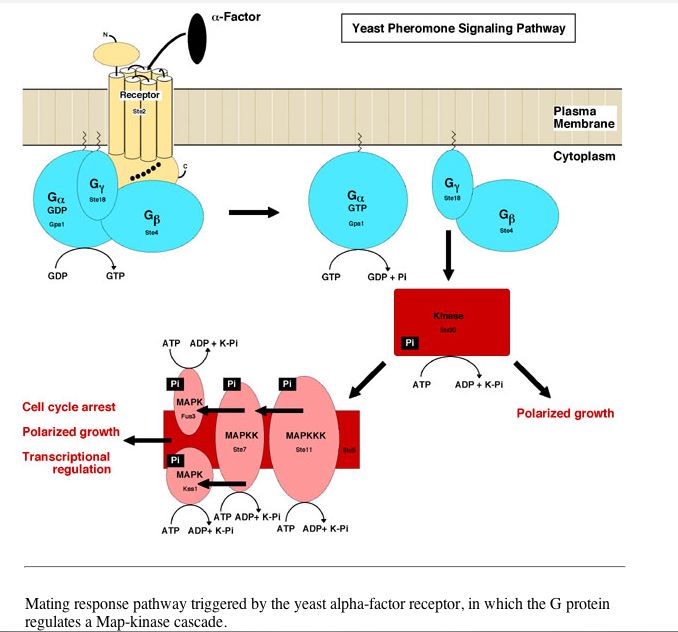
Image and caption provided by: http://spot.colorado.edu/~falke/figures/fig11/figure11.html
Expression Pattern 3: in response to alpha-factor (over time)
.jpg)
Fig. 5. Clustered expression of CYT1 to alpha-factor over time (SGD, 2004).
.jpg)
Fig. 6. CYT1 expression changes over time when exposed to alpha-factor (SGD, 2004).
The expression of this combination of clustered genes in response to alpha-factor over time appears repressed. It appears that repression increases for the first 45 minutes of alpha-factor exposure after which repression decreases. CYT1 appears not to be expressed as heavily in the presence of alpha-factor. Perhaps energy is conserved during yeast sexual reproduction, and CYT1 is down regulated.
Expression Pattern 4: in response to alpha-factor (various concentrations)
.jpg)
Fig. 7. Genes clustered with CYT1 according to expression correlation with various concentrations of alpha-factor (SGD, 2004).
.jpg)
Fig. 8. Expression of CYT1 changes over time in response to a varying concentration of alpha-factor (SGD, 2004).
The expression of CYT1 in response to various concentrations of alpha-factor is different from that of CYT1 in response to the amount of time CYT1 is exposed to alpha-factor. With varrying concentrations, the expression response is sometimes repressed and sometimes induced, but not in a linear dose-response manner. The expression response induction of CYT1 appears to peak around 200 nM. There could be points during the pheromone signaling pathway of alpha-factor at which the CYT1 electron transporter is needed, and is thus induced, and points during the same pathway at which CYT1 is not needed, and is therefore repressed.
Expression Pattern 5: in response to histone depletion
In this study, histone H4 depletion was caused by reducing nucleosome content and caused increased expression of 15% of genes, reduced expression of 10% of genes, and little effect on expression of the majority (75%) of yeast genes (Wyrick, 1999).
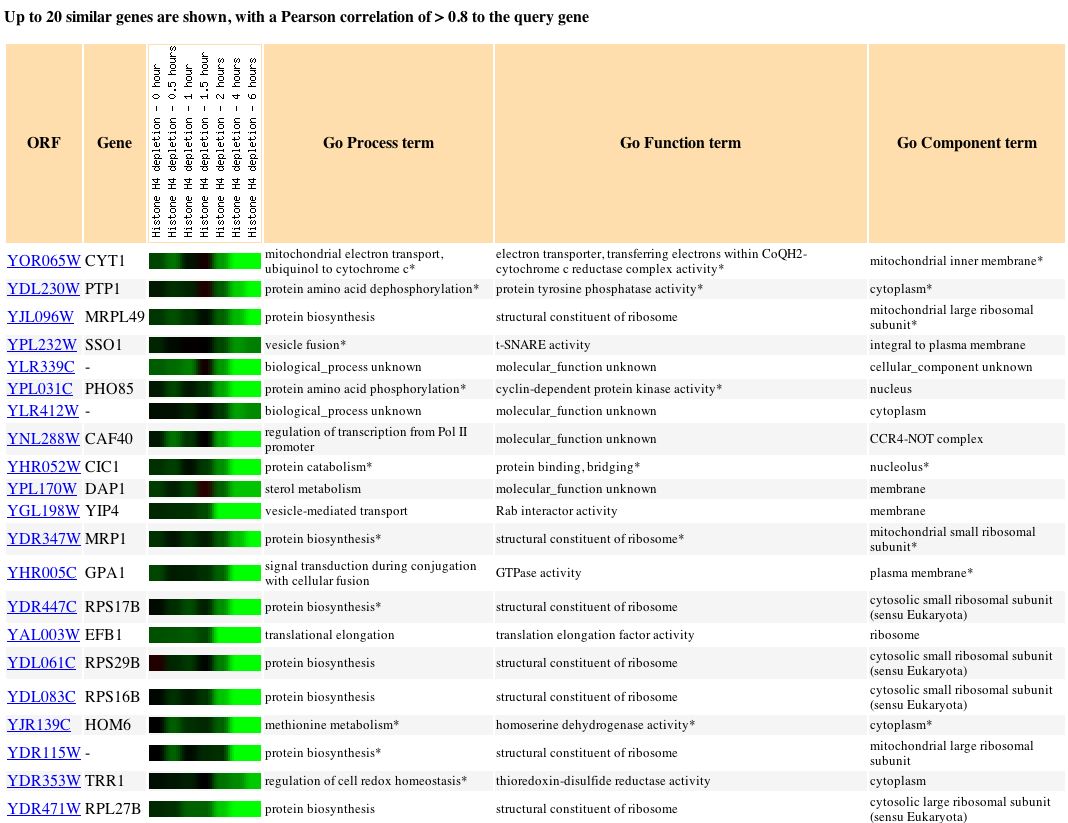
Fig. 9. Genes clustered according to similar expression patterns of histone depletion (SGD, 2004).
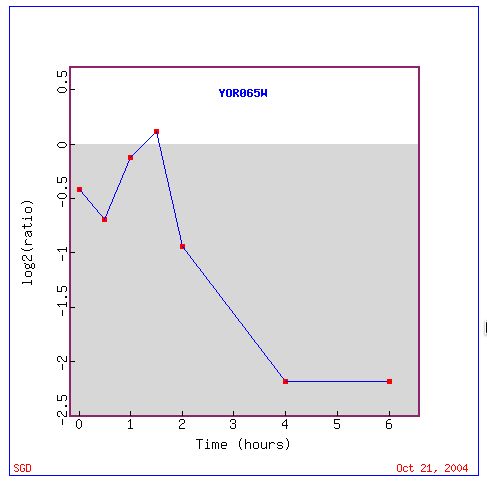
Fig. 10. Response of CYT1 to histone depletions over time (SGD, 2004).
Generally, histone depletion over time appears to repress CYT1 expression. However, because histones are located only in the nucleus, and CYT1 is located in the mitochondrial inner membrane, it is unlikely that histone depletion would have a direct affect on CYT1 expression. The data indicates that the role of histones may be gene specific rather than generally repressive.
Conclusions
After looking at DNA microarray clustered data and base 2 logarithm trend lines representing the microarray data for my favorite annotated gene, CYT1, it appears , based on the data regarding the diauxic shift, the alpha-factor responses and the response of CYT1 during sporulation that this gene plays a role in respiration and energy regulation or metabolism. These images alone would not tell the whole CYT1 story and there is a lot of information in these images that was not addressed. One helpful prospect of these data sets is the functional assignment of genes that currently have an unknown biological process but are clustered with the known gene, CYT1. Although it was not clustered in any of the above figures, we will now turn to expression patterns of my favorite non-annotated gene YOR066W, which is located near CYT1 on Saccharomyces cerevisiae, chromosome 15.YOR066W
In a previous web page, the gene YOR066W, a non-annotated gene in Saccharomyces cerevisiae, was also examined. Its molecular and cellular functions as well as its biological process are unknown. To better understand this gene, the same conditions examined above were analyzed using Expression Connection website. The spectrum below shows again the scale with which microarray clustered data was analyzed.
Scale: (fold repression : fold induction)

Expression Pattern 6: during sporulation
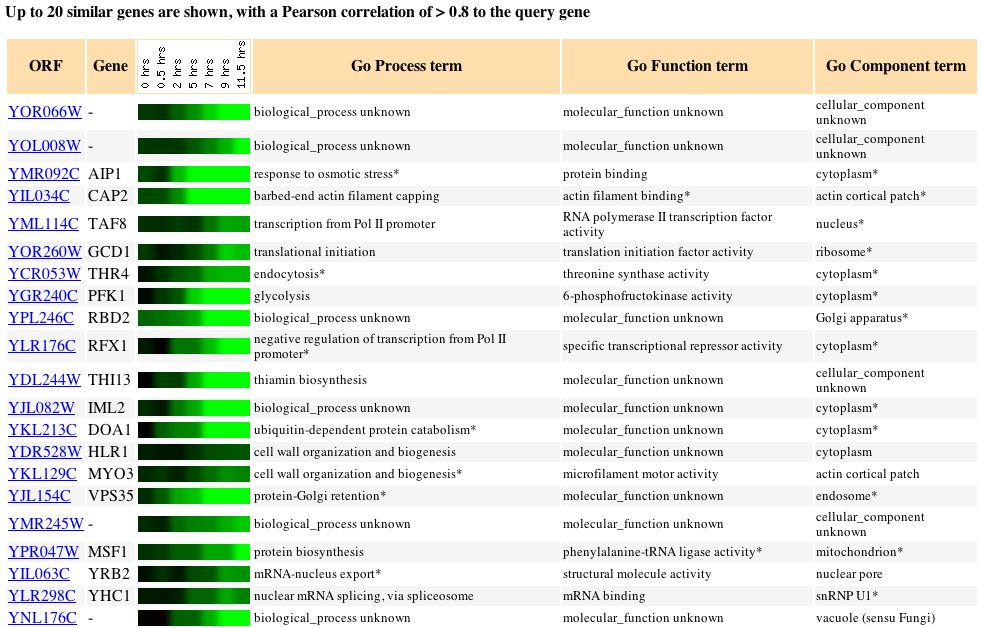
Fig. 11. Clustered expression pattern of YOR066W during sporulation (SGD, 2004).
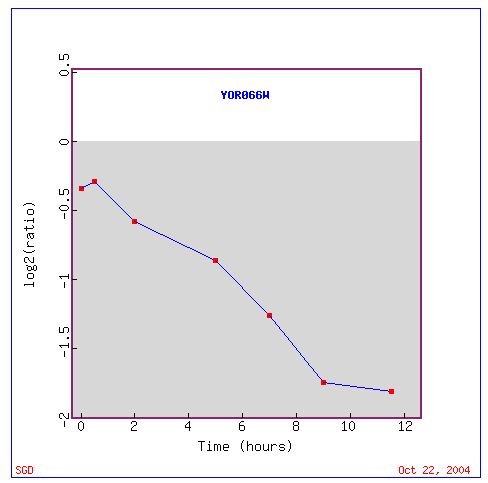
Fig. 12. Response of YOR066W to sporulation over time (SGD, 2004).
YOR066W is clustered with other genes that show repression during biogenesis and biosynthesis processes. YOR006W may still be involved in the cellular processes such as transcription and translation, as indicated by clustered genes, but YOR006W appears to be repressed during sporulation.
Expression Pattern 7: the diauxic shift
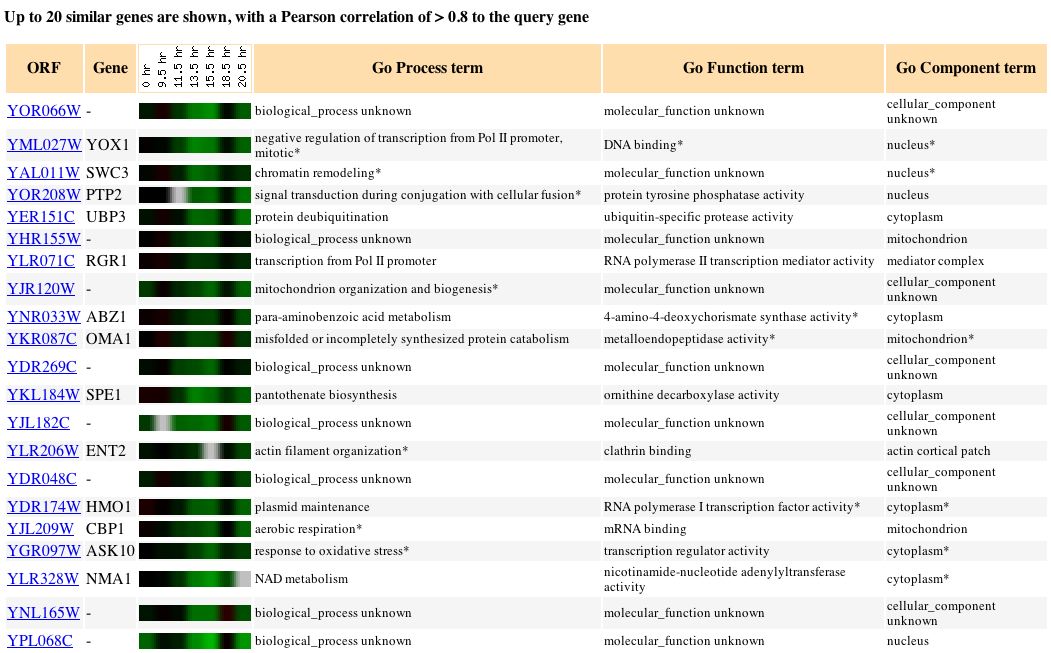

Fig. 13. Microarray data for YOR066W during the diauxic shift (SGD, 2004).
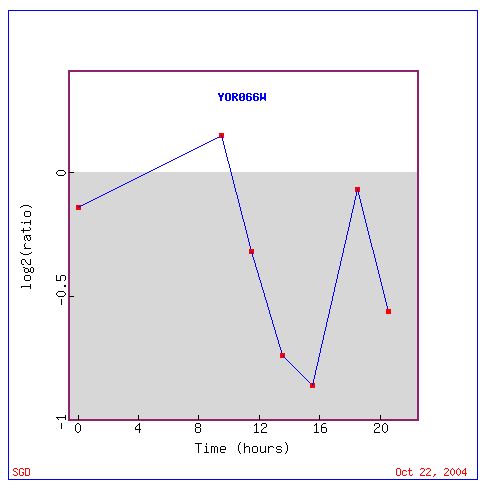
Fig. 14. Expression response for YOR066W during the diauxic shift (SGD, 2004).
This data is interesting because it is so different from the response of CYT1 under the same experimental conditions. This might mean that the two genes are completely unrelated. Another hypothesis is that the two genes are regulated similarly, but have opposite biological functions (i.e. CYT1 is turned on by a protein that turns YOR066W off under conditions of diauxic shift). Such is the case for yeast genes involved in the environmental stress reponse (ESR) (Campbell, 2004). Because diauxic shift is a "stressful" condition for yeast, it is possible that YOR066W is involved among thee 900 yeast genes, in the ESR.
Expression Pattern 8: in response to alpha-factor (over time)

Fig. 15. Clustered expression patterns of YOR066W when exposed to alpha-factor, over time (SGD, 2004).
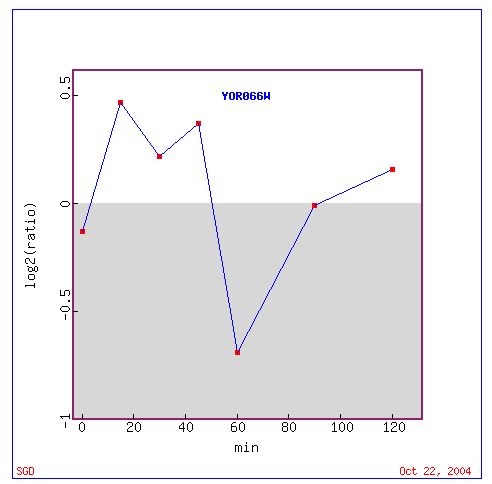
Fig. 16. Response of YOR66W to alpha-factor exposure, over time (SGD, 2004).
Fig.16 shows a similar response to that of CYT1 when exposed to alpha-factor, over time. It should be noted however that while a similar level of expression can be seen in the two experiments, the maximum repression of YOR66W occurs at a later time point.
Expression Pattern 9: in response to alpha-factor (various concentrations)

Fig. 17. Expression pattern of genes clustered with YOR066W at various alpha-factor concentrations (SGD, 2004).
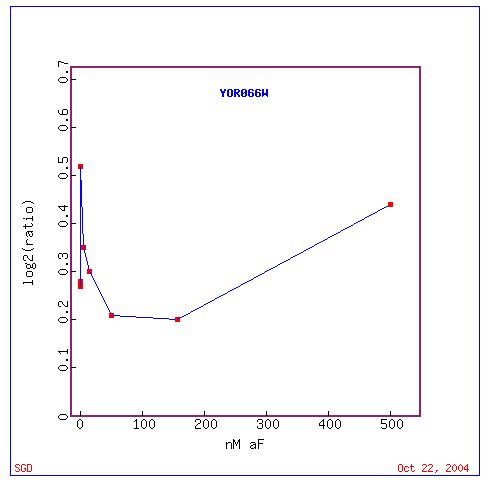
Fig. 18. Response of YOR066W at varied alpha-factor concentrations (SGD, 2004).
This is an interesting graph when we compare CYT1 response with YOR066W at varried alpha-concentrations. The two genes have distict points of similar base 2 logarithm ratios around concentration 200nM. However for CYT1, this is the maximum induction and for YOR0066W this is the point of minimum induction.
Expression Pattern 10: in response to histone depletion

Fig. 19. Genes clustered according to similar expression patterns of histone depletion (SGD, 2004).
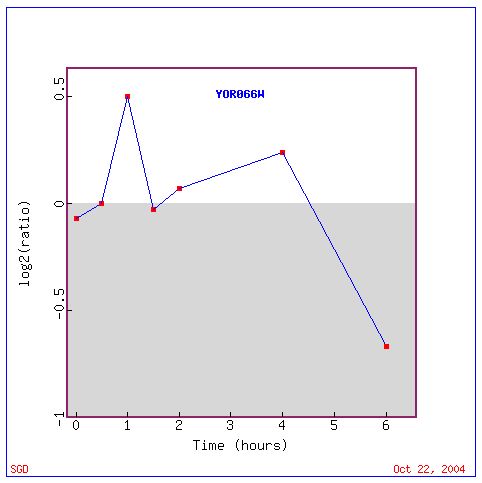
Fig. 20. The response of YOR066W to histone depletions over time (SGD, 2004).
The response of YOR066W to histone depletion shows a similar trend to that of CYT1. Looking at the clustered genes for this experiment, there is a possibility that this non-annotated gene is involved in chromatin remodeling or assembly, DNA repair and/or a number of other cellular roles indicated by established and non-annotated genes.
Conclusions
While a previous investigation, based on amino acid sequence similarity, suggested that YOR066W potentially coded for a CDC28 substrate, such data was not supported, here. A "CDC28 substrate" did not appear as a clustered gene in any of the microarray expression patterns. It is still possible that YOR066W codes for a CDC28 substrate that was represented by another Go function term in the clustered data. Generally, responses of YOR066W did not match those of CYT1. YOR066W responses to the tested conditions were similar to clustered gene responses. These cluster mates are protein function candidates. However, there was no one clustered gene that repeatedly appeared in microarray clusters for different experimental conditions. The only thing that can be concretely concluded about this yeast gene YOR066W from this investigation, is that more research must be done in order to fully understand its function.
References:
Balakrishnan R. SGD Currator. <yeast-curator@genome.stanford.edu> Personal communication. email. 2004 Oct 29.
Campbell MA, Heyer LJ (2003) Discovering genomics, proteomics, and bioinformatics. Bengamin Cummings: New York.
Diagram from DeRisi JL, Iyer VR, Brown PO (1997) Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278:680-686.
Wyrick JJ, Holstege FC, Jennings EG, Causton HC, Shore D, Grunstein M, Lander ES, Young RA (1999) Chromosomal landscape of nucleosome-dependent gene expression and silencing in yeast. Nature 402(6760):418-21.
SGD, Expression Connection. 2004. < http://db.yeastgenome.org/cgi-bin/expression/expressionConnection.pl> Accessed 2004 Oct 21.
DNA Microarray Methodology - Flash Animation, 2003. http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/method/chip/chip.html Accessed 2004 Oct 21.
Emily Wilson's Genomics Home Page
Email Questions, Comments or Suggestions : emwilson@davidson.edu