Figure 1: DMC1 predicted structure. Image provided by the Protein Data Bank.
*This web page was produced as an assignment for an undergraduate course at Davidson College.
Investigating the Gene Products of DMC1 and YER185W
Introduction
Previously, DMC1 and YER185W were examined on the levels of genes and gene expressions. These two ORFs will now be investigated on the gene product, or protein, level. Information regarding the structure and location of the protein helps investigators understand the diverse functions of proteins.
DMC1
First we will look at the structure of DMC1 (Figure 1). Notice how the individual subunits come together to form a circle but leave a hole in the middle. Such a shape is consistent with the predicted function of this gene product. It has been shown that DMC1 is involved in repairing single and double-stranded breaks in DNA following recombination. The circular shape is conducive to threading the DNA in need of repair through the protein as needed.
Figure 1: DMC1 predicted structure. Image provided by the Protein Data Bank.
We can then take a look at interactions of DMC1 with other proteins through Yeast Two Hybrid Data (Figure 2). In this method, the protein of interest, in this case DMC1, is the bait while other proteins suspected to interact with DMC1 are the prey. When the bait and prey interact, transcription of a reporter gene occurs. To learn more about the Yeast Two Hybrid method, visit the molecular biology animation. It seems that DMC1 (YER179W) interacted with 6 different proteins, one as the prey and five as the bait.
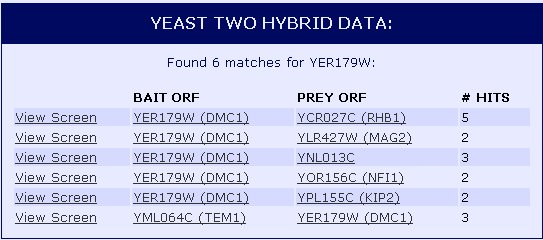
![]() Figure 2: Protein-protein interactions of DMC1 (YER179W) shown through Yeast Two Hybrid method. Data provided by the Yeast Resource Center.
Figure 2: Protein-protein interactions of DMC1 (YER179W) shown through Yeast Two Hybrid method. Data provided by the Yeast Resource Center.
The Yeast Grid database displays many other possible proteins with which DMC1 is likely to interact (Figure 3). DMC1 is believed to be involved in DNA repairs following recombination in meiosis; therefore, it follows that many of the proteins with which DMC1 interacts are involved in DNA binding during replication, transcription, and translation.
![]()
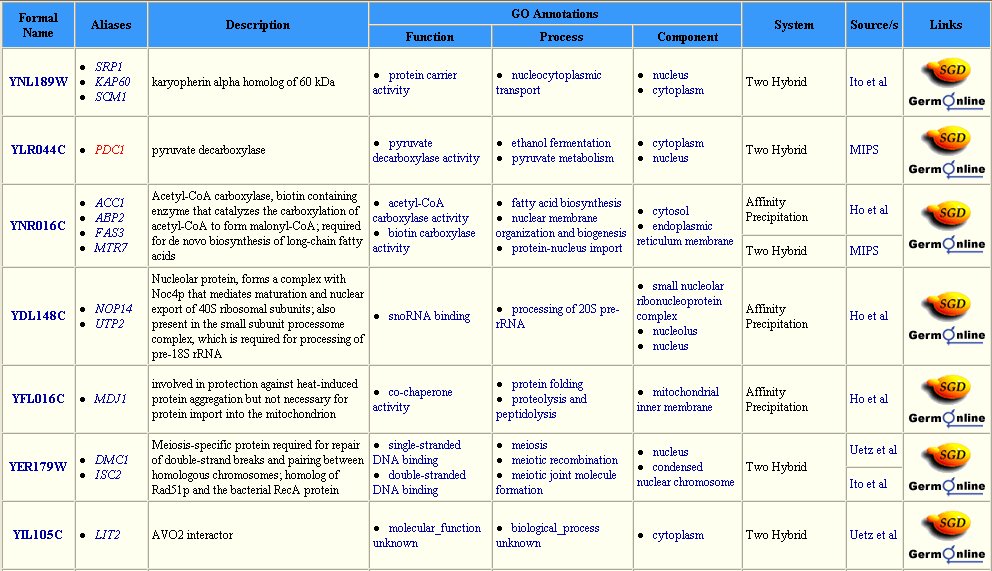
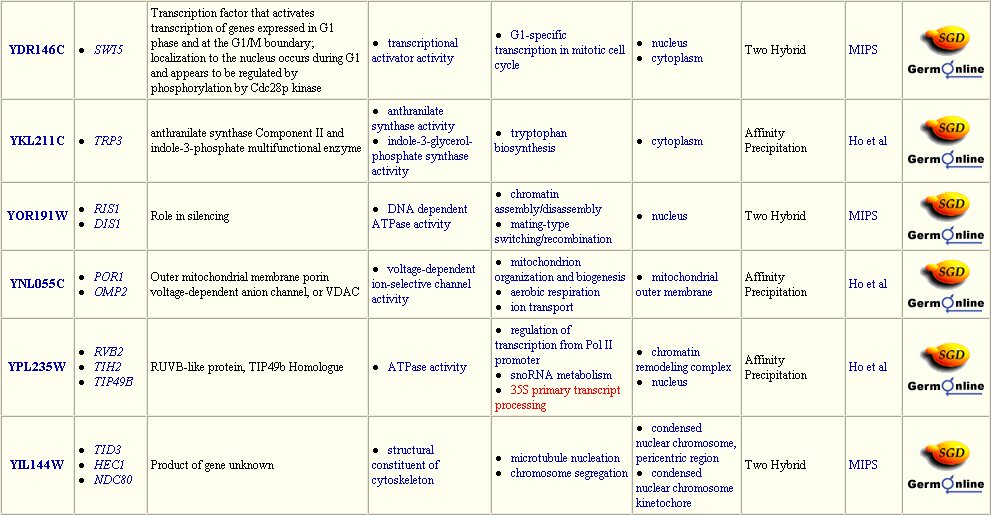
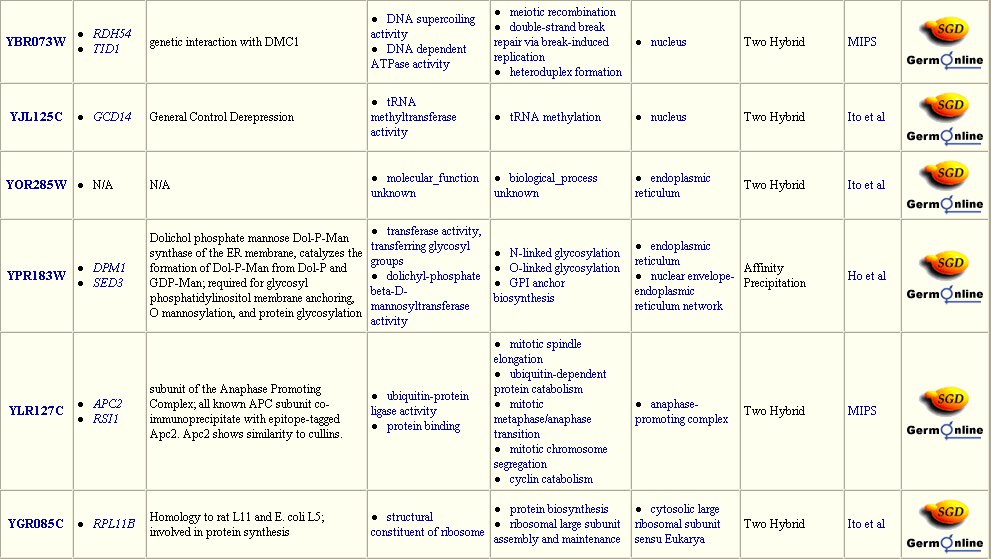
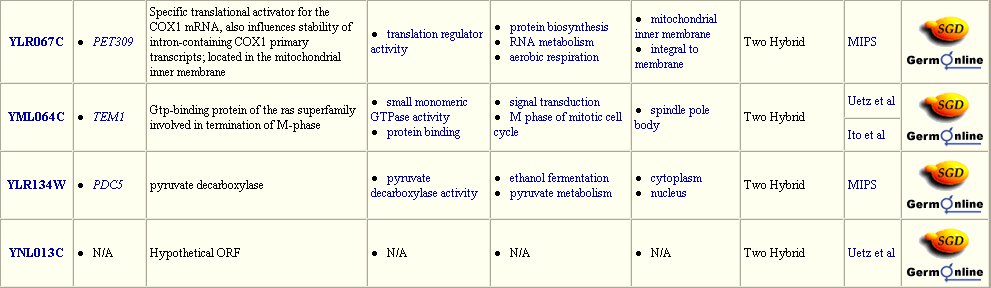
Figure 3: Interaction between DMC1 and each of these twenty-three proteins may occur. Many of these proteins are involved in DNA binding in transcription or translation. Data extracted from the Yeast Grid database.
The interactions of DMC1 with a small number of the proteins listed in the grid above have been verified and are included in the graph below (Figure 4). The central red dot is DMC1. The proteins most closely interacting with DMC1 are the dots directly connected to DMC1. Interactions verified by methods other than high-throughput methods are depicted with a green line and include: Srp1p, Tem1p, Apc2p, Acc1p. For further explanation of this graph, see the legend below (Figure 5).
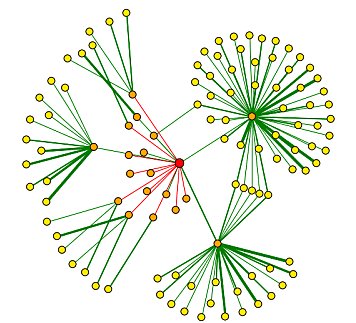
Figure 4: Graph of DMC1 protein-protein interactions. The red central dot represents DMC1. For further explanation of this graph, see the legend below (Figure 5). Image provided by the DIP database.
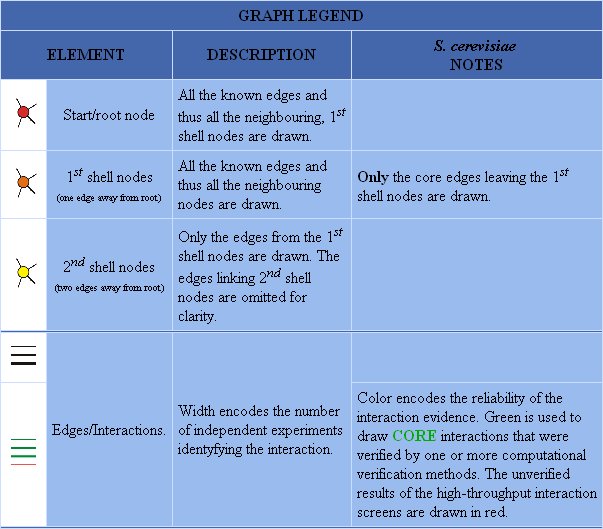
Figure 5: Graph legend for protein-protein interactions of DMC1.
The DIP database gives further information on the proteins that closely interact with DMC1 (Table 1). All of these proteins have a role in the nucleus or during meiosis. Therefore, the fuctions of these proteins are consistent with prior knowledge of DMC1.
Table 1. GO function information concerning DMC1 and proteins with which it interacts.
| Name | Process | Function | Localization |
| DMC1 | meiotic recombination meiotic joint molecule formation |
single-stranded DNA binding double-stranded DNA binding |
condensed nuclear chromasome nucleus |
| Srp1p | nucleocytoplasmic transport | protein carrier activity | cytoplasm nucleus |
| Tem1p | M phase of meiotic cell cycle signal transduction |
protein binding GTPase activity |
spindle pole body |
| Apc2p | hypothetical protein | unknown | unknown |
| Acc1p | fatty acid biosynthesis nuclear membrane organization and biogenesis protein-nucleus transport |
biotin carboxylase activity acetyl-CoA carboxylase activity |
ER membrane mitochondria cytoplasm |
YER185W
There is much less information pertaining to the gene product of YER185W. For example, while there were twenty-three possible proteins listed in the Yeast Grid database that interact with DMC1, there are none listed for YER185W (Figure 6).
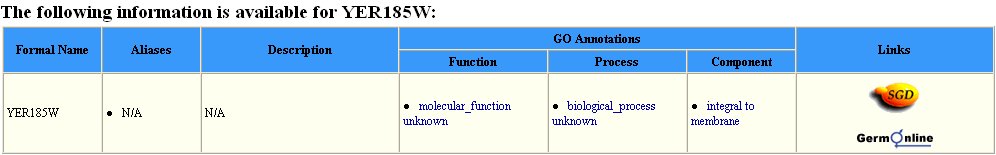 Figure 6: Search results in the Yeast Grid database for YER185W. Much less information is known about this protein than is known of DMC1.
Figure 6: Search results in the Yeast Grid database for YER185W. Much less information is known about this protein than is known of DMC1.
The KEGG database gives more data, however. There is no pathway information available for YER185W, but there is data describing orthologs, paralogs, and motifs of this protein. A graph of clustered orthologs is made for YER185 and visualized through the OC Viewer (Figure 7).
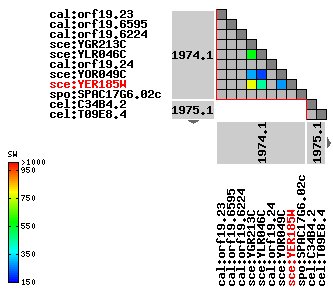
Figure 7: Ortholog clusters for YER185W. YER185W clusters closely with YGR213C. Image is reproduced from the KEGG OC Viewer.
YER185W contains the conserved domain, RTA1, shared by the proteins below (Figure 8). Especially interesting is the protein YGR213C. We saw above that this protein clusters closely with YER185W. Below we can see that it also contains the RTA1 domain. YGR213C is involved in 7-aminocholesterol resistance.

Figure 8: Descriptions of proteins sharing the RTA1 conserved motif (data from KEGG).
The KEGG website also lists many orthologs for YER185W (Figure 9). An ortholog is an analagous gene in a different species. However, the most similar ortholog has only a 0.272 identity with YER185W. Therefore, the functions of the gene products of these orthologs are probably not as indicative as they could be of the function of YER185W.
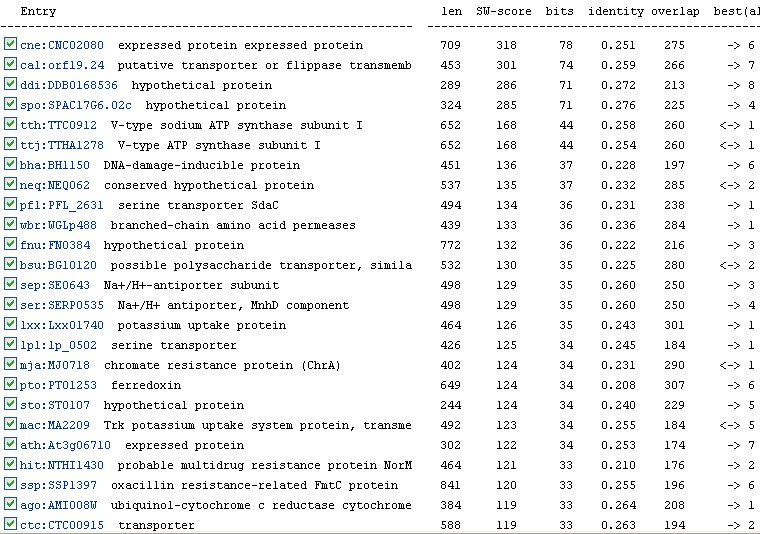
Figure 9: Orthologs of YER185W as listed by the KEGG database.
In addition to orthologs, the KEGG database also lists six paralogs for YER185W (Figure 10). A paralog is a duplicated gene in the same organism that has evolved away from the original gene. The identities of the paralogs are slightly better than those of the orthologs. Again, we see the protein YGR213C listed as the most similar to YER185W.
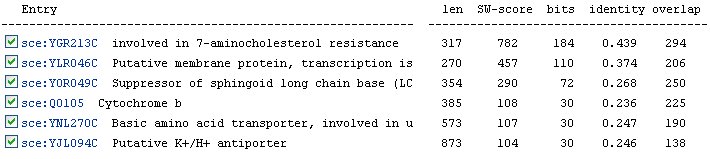 Figure 10: Paralogs of YER185W as listed by the KEGG database.
Figure 10: Paralogs of YER185W as listed by the KEGG database.
The Protein Information Resource (PIR) offers some information concerning YER185W, but this information is not much more specific than the information found in expression data. It is found through this website, however, that this protein exhibits a response to xenobiotic stimuli (Figure 11). This is consistent with the fact that YER185W contains a conserved domain held by other proteins aiding in antibacterial resistance.
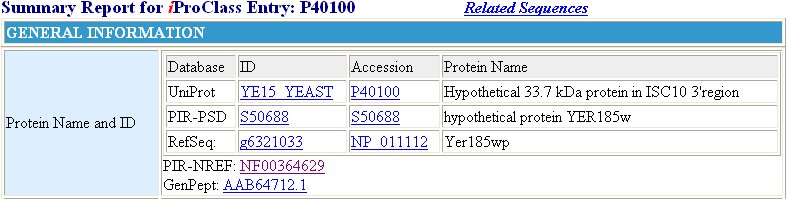

Figure 11 : Information provided by the PIR database for YER185W. We find from this database, again, that YER185W is a transmembrane protein. Also learned from this information is the fact that this protein has a molecular weight of 33.7 kDa, and it does respond to xenobiotic stimulus.
Yeast Two Hybrid experiments performed on YER185W only show interaction with one protein (Figure 12). This protein, YJL147C is also uncharacterized. The cellular component of YJL147C is thought to be the mitochondria, but little else is known about this protein (SGD).

Figure 12: Results of Yeast Two Hybrid experiments for YER185W. YER185W interacts with possibly one protein, YJL147C. Data taken from the Yeast Resource Center .
Another way to extract information about a protein is to disrupt the protein and see what the effect is. This is what was done in the TRIPLES database. In YER185W, deviations occurred from wild type in the case of the HapTra assay and the 003SDS assay (Figure 13). A weak deviation in the HapTra assay indicates that the gene may be haplosufficient, but there is a decreased efficiency of the protein. A strong deviation from wild type in the 003SDS assay indicates strongly that YER185W is involved in cell wall maintenence and/or biogenesis.
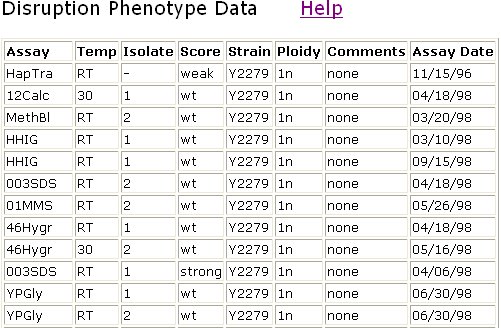
Figure 13: Phenotypes were scored after disrupting the gene encoding YER185W. A change in HapTra indicates that haploid mutants are inviable. Changes in 003SDS indicates that a gene may encode a protein involved in cell wall maintenance and biogenesis. The score column indicates the deviation from wild type. A score of "wt" indicates no significant change from wild type. Deviations from wild type reactions to these assays are scored on a range from weak to strong. Data reproduced from the TRIPLES database.
Conclusion
The data for DMC1 overwhelmingly show that it is a protein involved in DNA repair after crossing over in meiosis. This protein's shape and interactions with other proteins show that it is involved in repairing both single-strand and double-strand breaks in DNA. This is further supported by the fact that this protein has been localized to the nucleus.
The data support the theory that YER185W is a transmembrane protein. It is likely that this protein aids in conferring some type of antibiotic resistance to the organism. This inference is drawn from the orthologs, specifically YGR213C, with which YER185W clusters. There are also data supporting the idea that YER185W is involved in cell wall maintenance and biogenesis.
Future Experiments
Considering what we know about YER185W, and more importantly, what we do not know, I would suggest that the first wet lab test that be done would be an immunoprecipitation. It would be helpful to see better with which, if any, other proteins YER185W co-precipitates. This procedure is more reliable than the yeast two hybrid method because it does not involve an artificial construct. To do this, an antibody would have to be raised against this protein and characterized. While this is a somewhat complicated process, the advantages of having an antibody against a protein of interest are numerous.
I would also suggest that the cellular location be found through one of many methods. It is assumed that this protein resides in a membrane. To determine whether this is the plasma membrane or the membrane of an organelle, a mild detergent could be used to rupture the outer membrane but not the inner membranes. Of course, the cell wall would have to be breached as well. After this, if YER185 is in the plasma membrane, it will now be free-floating and able to locate. If it is still tied up in a membrane, it will spin down with a pellet when centrifuged and will not be found in the supernatant mixture.
To localize the protein immunofluorescence could also be used, especially if an anti-YER185W antibody is already being created for the immunoprecipitation..
Also, because a current theory is that YER185W plays a part in antibacterial resistance, studies could be done in the presence of various common antibotics. In these studies, yeast with a null mutation in this gene, yeast with one good copy of the gene, and yeast with two good copies of the gene could be compared.
Works Cited:
KEGG database.
Kumar, A., Cheung, K.-H., Ross-Macdonald, P., Coelho, P.S.R., Miller, P., and Snyder, M. (2000). TRIPLES: a Database of Gene Function in S. cerevisiae. Nucleic Acids Res. 28, 81-84. (Full-text in PDF reproduced with permission from NAR Online http://www.oup.co.uk/nar )
Protein Information Resource (PIR)
Ross-Macdonald, P., Coelho, P.S.R., Roemer, T., Agarwal, S., Kumar, A., Jansen, R., Cheung, K.-H., Sheehan, A., Symoniatis, D., Umansky, L., Heidtman, M., Nelson, K., Iwasaki, H., Hager, K., Gerstein, M., Miller, P., Roeder, G.S., and Snyder, M. (1999). Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402, 413-418. Nature 402, 413-418.
Saccharomyces Genome Database (SGD)
If you have any questions or concerns about this webpage please contact Sarah.
Return to Davidson College Biology Home Page.
Return to Genomics Home Page.
Return to Sarah's Home Page