The Ernest Gallo Clinic and Research Center in the University of California at San Francisco’s Department of Neurology has found a gene responsible for the effects of ethanol on the model organism Caenorhabditis elegans. Read on to get a feel for how the popular press is portraying the discovery, and then learn more about the actual experiment by reading an in-depth review of the scientific paper.
I. The Popular Press on “The Intoxication Gene”
It was only to be expected that, when Stephen McIntire and others in his lab at the University of California in San Francisco doused worms with alcohol to test a gene that they suspected mediates intoxication, the popular media would have a few quips to make. In one MSNBC story entitled “'Intoxicating' gene found in worms,” the experimental groups are jokingly referred to as “soused” or “teetotalers.” But jokes aside, how accurately and effectively does the popular media report scientific findings?
Kua et al. (2004), in their paper entitled “Science in the news: a study of reporting genomics,” address the issue of popular media’s science coverage, saying “articles frequently [omit] methodological and contextual information, features most often mentioned as critical for a complete journalistic account of science” (Kua et al. 2004). They go on to present tools a science writer should provide his or her readers, namely “a good explanation of the science, […] questions about the long-term significance of the work, […] and giving direct context [including] the current body of knowledge in the subject and the state and direction of research in the field” (Kua et al. 2004).
I have found two articles on “the Intoxication gene,” and both of them seem to fall short when judged by Kua et al.’s description of the science writer as a “tool-giver.” Let’s give them each a look:
In MSNBC’s Associated Press story, “'Intoxicating' gene found in worms,” the experiment is depicted in terms of its long-term significance, focusing on McIntire’s hope that they will eventually “find a way to cure alcoholism and drug abuse” (MSNBC). The author even introduces some of McIntire’s methodology, explaining that worms were exposed to alcohol and those who had a particular mutated gene (which remains unnamed in the article) were found to be resistant to the effects of ethanol. However, the author fails to place the experiment in the context of other research. Davies et al. (2003) reveals in his paper that other ethanol-response pathways are being explored using their methods of mutagenesis to find genes implicated in ethanol sensitivity. This information is crucial to communicating to the reader that science is a “puzzle-solving process” and that pieces are constantly being found and connected (Kua et al. 2004).
Later, the author says “Alcohol increases the gene’s activity, which slows down brain activity […]. But if the gene is disabled, as it was in the mutant worms, the brain never gets the chance to slow down. Still, McIntire [cautions that] there’s much research left to do before the leap to people can be made” (MSNBC). The obvious question here is “what leap?” The author never reveals the significance of the mutated gene’s lack of function: namely, that in humans with defective orthologs of this gene, a higher volume of alcohol will have to be consumed to produce the same effects, thereby increasing the risk of alcoholism. This omission leaves the readers in the dark about the true nature of the “intoxication gene” and potential treatments for alcoholism.
The second article was found on the Health on the Net (HON) Foundation’s web site, and is titled “Gene That Causes Alcohol Intoxication Found” by Steven Reinberg. Right away, the title seems to commit a common media error described by Marianna Pellechia and quoted in Kua et al. (2004); that is, an “omission of qualifying statements.” In actuality, the gene that researchers found was only one of the genes that play roles in alcohol intoxication. Indeed, Davies et al. (2003) is careful to caution that they “think it is unlikely that [the gene’s target is] the only ethanol target.” However, in the first two paragraphs of the HON article, the same oversimplification is repeated twice. In the latter part of the article, Reinberg goes so far as to create the idea of a scientific debate about whether McIntire’s “intoxication gene” is the only gene involved in intoxication—something McIntire does not actually assert in his paper. Reinberg writes “While [scientist] Treistman agrees [with McIntire that] the BK channel has to be a target in controlling the effects of alcohol, he says it's not the only target” (Reinberg).
The misguided slant of the article continues when Reinberg says “Basically, McIntire says, if you take away the intoxication, you take away the desire and hence the dependence” (Reinberg). However, this is the opposite of what McIntire states in his paper, namely that “In humans, variation in behavioral sensitivity to ethanol has […] been found to correlate with frequency of alcoholism—individuals with lower sensitivity to ethanol have a higher frequency of alcoholism” (Davies et al. 2003, emphasis added). In other words, McIntire is stating that the mutation of the “intoxication gene,” which lessens the intoxication, is a risk factor for alcoholism—not that lessening the intoxication lowers the risk of alcoholism.
The HON article does have some redeeming qualities. It briefly mentions methodology and the context of the discovery, noting that “other genes have been identified that are very important in […] turning on drug-related behavior” (Reinberg). It also remarks on the long-term potential of the study to play a role in curing alcoholism.
As a whole, however, both of these articles fall short in reporting this scientific discovery of an ethanol response pathway. While they both delve into long-term significance, presumably to hook their audience with how this story may relate to them, they provide only brief methodology of the experiment and fail miserably to place the study in the context of current research.
Now let’s look at the actual study to see what we’ve missed reading the popular press articles.
II. The Scientific Scoop on slo-1
The human behavioral responses to ethanol have been well-documented. Loss of coordination and social inhibition, incoherency, and sedation are all symptoms of intoxication (Davies et al. 2003). However, the mechanisms that cause the behavioral changes due to intoxication are not well-understood. Thus, Davies et al. (2003) performed an experiment designed to elucidate the molecular mechanisms that underlie the behavioral effects of intoxication. They used the model organism C. elegans, which becomes intoxicated at the same dosage of ethanol that intoxicates humans.
There are two tactics that can be used to study the molecular mechanisms of intoxication. The first is a biochemical method in which specific ethanol targets are sought. At least 20 such target proteins have been found via this method, but their role in intoxication is not yet known (Davies et al. 2003).
The second method, and the one employed by Davies et al. (2003), is a behavioral analysis of mutant model organisms exposed to ethanol. The premise of this method is that, if researchers find mutants whose behavior does not appear to be influenced by ethanol, the common gene these mutants lack may play a role in the intoxication behaviors of wild-type individuals. To utilize this method, there must be distinct intoxication behaviors in the model organism by which ethanol sensitivity can be measured. For C. elegans, the intoxication behaviors are decreases in speed of locomotion, amplitude of body waves as they crawl, and egg-laying frequency (Figure 1).
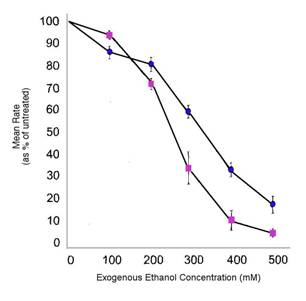
Figure 1. Effects of ethanol on C. elegans rate of locomotion and egg-laying. Both rates dramatically decrease in wild-type worms in the presence of ethanol. Locomotion speed is shown by blue circles while egg-laying frequency is shown by purple squares. Adapted from Davies et al. 2003.
This method has been successfully used in past research to find genes that affect sensitivity to ethanol. For instance, the gene for neuronal nitric oxide synthase (nNOS) was knocked out in experimental mice so that a behavioral assay could be conducted to determine nNOS’ role in ethanol sensitivity (Spanagel et al. 2002). The nNOS knockout mice (nNOS -/-) and the wild-type mice were then given a choice between two bottles, one with water and the other with increasing concentrations of alcohol. When the alcohol was offered at low concentrations, there was no significant difference between the choices of the nNOS -/- mice and the wild-type mice. However, when alcohol was offered at higher concentrations, the knock-out mice drank six times as much alcohol as the wild-type mice. The nNOS -/- mice were less sensitive to ethanol as measured by their quicker recovery from the sedative effects of ethanol. By conducting this behavioral assay on the mutant strain of mice, Spanagel et al. (2002) were able to find a gene that plays a role in ethanol tolerance and thus alcoholism. Other genes have also been found to play a role in ethanol’s effects using the behavioral analysis method (Table 1).
Table 1. Several genes found to alter ethanol sensitivity in behavioral assays. By observing the behavior of mutant organisms in response to ethanol, it is possible to find genes that play a role in intoxication.
Researchers exposed wild-type worms to ethane methyl sulfonate ( EMS), a chemical known to cause genetic mutations, for 4 hours. This process is called mutagenesis and creates a population of mutants, or organisms with different genetic profiles than that of a wild-type individual. They then screened the mutants for resistance to ethanol. To do this, they completed both a locomotion and an egg-laying assay. For the locomotion assay, they incubated mutants in 400 mM ethanol, put them in the presence of a substance with an attractive odor, and selected mutants that showed no significant decrease in speed or amplitude of body waves. In the egg-laying assay, they exposed mutants to 500 mM of ethanol, and selected those that could still lay eggs.
After selecting the resistant mutants, they had to determine the genetic map location of the worms’ mutation. It mapped to Chromosome V, where the slo-1 gene is located (Figure 2)
Figure 2. A representation of C. elegans’ chromosome V. A red arrow shows the location of the slo-1 gene. The gene covers 15.89 kb and contains 24 introns. More views of the C. elegans’ genome are available at the NCBI MapViewer. More information about this gene can be found at AceView.
slo-1 is named after a Drosophila ortholog called slowpoke. It also has orthologs in Homo sapiens (humans), Mus musculus (mice), and Gallus gallus (chickens) as well as other species. slo-1 codes for a calcium-activated BK potassium channel. Ethanol causes an increase in the BK potassium channel’s current, which inhibits neuron activity (when loss-of-function slo-1 mutants’ neuronal currents were measured, they were missing the BK potassium channel current that was present in the wild-type worms’ neurons). As aforementioned, a loss of slo-1 function caused C. elegans mutants to show resistance to the effects of ethanol (Figure 3). In contrast, hyperactivation of the BK potassium channel causes C. elegans to mimic intoxicated behavior.
Figure 3. Worms who have lost slo-1 function are resistant to the behavioral effects of ethanol. (A) Locomotion rates of wild-type worms exposed to ethanol compared to resistant slo-1 mutants exposed to ethanol (+/- SEM) (B) Egg-laying frequency of wild-type worms exposed to ethanol compared to resistant slo-1 mutants exposed to ethanol (+/- SEM). Asterisk indicates significance of P < 0.01. Adapted from Davies et al. 2003.
Treatment |
Link to Movie |
untreated wild-type | |
wild-type exposed to ethanol | |
resistant exposed to ethanol |
Table 2. Movies depicting C. elegans behavior in response to various treatments. Quicktime is required to view these movies (The Health Report).
After seeing the effects of both the loss of slo-1 function and the overexpression of slo-1, the researchers felt confident that slo-1 was responsible for controlling “acute neurobehavioral effects of the [ethanol]” (Davies et al. 2003). However, there were still several more questions that needed to be asked in order to clarify slo-1’s role in intoxication.
d. Asking More Questions about slo-1
- Is the ethanol resistance shown by slo-1 mutants merely due to changes in ethanol metabolism or actually due to changes in the ethanol response?
| To ensure that slo-1 mutants’ lack of ethanol-induced behavior was due to changes in the actual ethanol response and not merely to changes in how much ethanol was absorbed, researchers measured the internal concentration of ethanol in both the mutants and the wild-type worms. There was no significant difference in the internal concentrations of ethanol in mutant versus wild-type worms, indicating that the lack of ethanol-induced behavior in mutants was, in fact, due to a change in the response and not merely the amount of ethanol absorbed. |
- Since slo-1 is expressed in both neurons and muscles, which cells—neurons or muscle cells—are causing the behavioral response to ethanol?
| Davies et al. transformed some of the slo-1 mutants using tissue-specific gene promoters to restore expression of slo-1 to either the neurons or the muscle cells. If slo-1 expression is restored in the neurons, ethanol sensitivity is regained. However, if slo-1 expression is restored in the muscle cells, the insensitivity to ethanol remains. Thus the expression of slo-1 in neurons is what plays a role in ethanol response. |
e. So what? What’s the Big Deal About Drunken Worms?
Since BK potassium channels are conserved in mammals, it is possible that these channels play an important role in human intoxication. It has been shown that people with lower sensitivity to alcohol have a higher frequency of becoming alcoholic, presumably because they have to drink more alcohol to experience the same effects (Schuckit 1994). Because of this relationship between ethanol sensitivity and alcoholism, understanding the molecular mechanisms that determine ethanol sensitivity may provide clues to the treatment of the disease.
One must be cautious about imagining the connection between slo-1 and ethanol sensitivity is the whole story. As aforementioned, previous studies have found numerous ethanol targets and it is still unclear how these targets function in intoxication. Other genes affecting the BK potassium channels and other pathways involved in the ethanol response may be important to the treatment of alcoholism as well.
III. Conclusions about Popular Press "science news" versus Scientific Literature
The writers of the two popular media reviews of McIntire’s slo-1 study fulfilled their role as mere “intermediary”—they translated a scientific paper into language with which the public is familiar (Kua et al. 2004). They at least attempted to satisfy the role of “watchdog”—discussing long-term implications of the study for the public (Kua et al. 2004). However, they falter when evaluated as a “tool-giver.” Tool-givers are writers who “give readers the tools with which to think and evaluate the evidence and the issues for themselves” (Kua et al. 2003).
The author of these two articles, touched only briefly on the methodology of the experiment and almost totally avoided the scientific context that surrounded the study’s findings. The public reading these articles will surely not understand the exciting process of finding slo-1 and determining its role in ethanol response. They will not have gained any knowledge about the research currently being conducted on the molecular mechanisms of drug response. Indeed, from at least one of the articles, they may get an altogether incorrect view of the significance of the findings. This humbling glance into the popular media’s portrayal of scientific findings suggests that there is a lot of work to be done on effective communication between scientists and the public.
Smith K.N., Iwanejko L., Loeillet S., Fabre F., and A. Nicolas. (1999) Disruption and functional analysis of seven ORFs on chromosome IV: YDL057w, YDL012c, YDL010w, YDL009c, YDL008w (APC11), YDL005c (MED2) and YDL003w (MCD1). Yeast 15(12): 1255-67 .
‘Intoxicating’ Gene Found in Worms. MSNBC. 11 Dec. 2003. 13 Sept. 2005. <http://msnbc.msn.com/id/3687545/>.
Kua, E., Reder, M., and M.J. Grossel. (2004) Science in the news: a study of reporting genomics. Public Understand. Sci. 13: 309-322.
Moore, M.S., DeZazzo, J., Luk, A.Y., Tully, T., Singh, C.M., and U. Heberlein. (1998) Ethanol intoxication in Drosophila: Genetic and pharmacological evidence for regulation by the cAMP signaling pathway. Cell 93(6): 997-1007.
Phillips, T.J., Brown, K.J., Burkhart-Kasch, S., Wenger, C.D., Kelly, M.A., Rubinstein, M., Grandy, D.K., and M.J. Low. (1998) Alcohol preference and sensitivity are markedly reduced in mice lacking dopamine D2 receptors. Nat. Neurosci. 1(7): 610-5.
Reinberg, S. “Gene That Causes Alcohol Intoxication Found: Discovery may lead to new treatments for alcohol abuse, researchers say.” Health On the Net Foundation. 12 Dec. 2003. 13 Sept. 2005. <http://www.hon.ch/News/HSN/516488.html>.
Rubinstein, M., Phillips, T.J., Bunzow, J.R., Falzone, T.L., Dziewczapolski, G., Zhang, G., Fang, Y., Larson, J.L., McDougall, J.A., Chester, J.A., Saez, C., Pugsley, T.A., Gershanik, O., Low, M.J., and D.K. Grandy. (1997) Mice lacking dopamine D4 receptors are supersensitive to ethanol, cocaine, and methamphetamine. Cell 90(6): 991-1001.
Shuckit, M.A. (1994) Low level of response to alcohol as a predictor of future alcoholism. Am. J. Psychiatry 151: 184-189.
Thiele, T.E., Marsh, D.J., Ste Marie, L., Bernstein, I.L., and R.D. Palmiter. (1998) Ethanol consumption and resistance are inversely related to neuropeptide Y levels. Nature 396(6709): 366-9.