A Review of "Synthetic Gene Networks That Count"
Introduction
Synthetic biology uses many tools from other disciplines, or seeks to imitate those tools within biological systems. The researchers in this case wanted to create a synthetic gene network that could be "programmed" to count to two or three like digital circuits can. Possible applications of this ability are being able to program cell death after a certain number of events, or knowing how many times a certain cellular event occurs. The researchers designed and built counters using two different methods: using regulation of the ability of the ribosome to bind to mRNA, and using enzymes that recombine DNA in a known way.
Riboregulated Transcriptional Cascade (RTC) Counters
Method
The counter constructs were created on plasmids that was then inserted into a strain of E. Coli. The two-counters and three-counters were built with the same general method, differing only in how many riboregulated units were involved in the counter set-up, the two-counter had 2 and the three-counter had 3 units. The general unit had, in order, a promoter, a cis-repressor (cr) sequence, a ribosome-binding site (RBS), and a gene. The cr sequence and the RBS were complentary, so when the unit was transcribed, a loop would form as the cr sequence and RBS bound together, blocking the ribosome from landing on the RBS and starting translation. This loop could be undone by a taRNA binding to the cr, freeing the RBS for the ribosome to bind to, starting translation.
Using these facts, the three-counter was designed to work like this:
(1) The cell starts with cr-RBS-T7 RNAP transcripts in it
(2) First pulse of arabinose binds to BAD promoter which transcribes the taRNA
(3) taRNA binds to cr of transcripts, allowing the ribosome to bind to RBS and express T7 RNAP
(4)Pulse ends, arabinose and taRNA are degraded, and T7 RNAP expression ends
(5) Already translated T7 RNAP transcribes cr-RBS-T2 RNAP
(6) Second arabinose pulse creates more taRNA
(7) taRNA binds to cr and frees RBS for ribosome binding and T3 RNAP expression
(8) Pulse ends, arabinose and taRNA are degraded, and T3 RNAP expression ends
(9) T3 RNAP already translated transcribes cr-RBS-GFP
(10) Third pulse of arabinose induces more taRNA
(11) taRNA binds to cr, freeing RBS for ribosome binding, and GFP is expressed
The end result of this design should be a cell that does not fluoresce until after the 3rd arabinose pulse The two-counter uses the same design only without the middle, T3 RNAP step, thus requiring only 2 pulses to express GFP.
Results
The experimental results supported the conclusion that the counter design worked. Cells not given any arabinose did not show any increase in mean fluorescence level. Those given less then a the full amount, whether one pulse for the two-counters or either one or two pulses for the three-counters, showed some increase in fluorescence, but none matched the increase that occurred in cells that received full dosages of arabinose. The limited increase in fluorescence for those cells not getting full doses was explained by the intended protein being expressed, but the cell also expressing a few proteins father down the chain than intended, resulting in fluorescence before it should have occurred.
Model Counters
Methods & Results
Using the design of their RTC counters, the authors built a mathematical model of the counters. They then used these models to make "predictions" for what the experimental results should have been, and analyzed how closely they matched the actual experimental results. The predicted results from the model closely matched those of the real experiments.
Once they authors were confident the model effectively represented what would actually occur, they used it to test the effects of varying the pulse duration and time between pulses. These expected results were also checked against actual results from experiments that varied the pulse duration and gap between them, and again the model closely matched the real-world results. This allowed the researchers to find the maximum and minimum pulse length and frequency, beyond which the counter could not correctly count due to kinetic limits of the cellular processes involved.
DNA Invertase Cascade (DIC) Counters
Method
The DIC counters were built using well-known and understood DNA recombinases Cre and flpe built into units called single invertase memory modules (SIMMs). Stringing together different numbers of these SIMMs on a plasmid allow the cells to count to either two or three depending on how many SIMMs the plasmid includes. The design of the three-counter is 2 SIMMs followed by the GFP gene. The 1st SIMM is the gene for flpe recombinase led by an inverted arabinose-activated promoter, and flanked by sites where flpe can cut, invert the DNA segment, and reattach in the new orientation. The 2nd SIMM is similar except being the cre gene and being flanked by the restriction sites for Cre, but with the same inverted arabinose-activated promoter. The last unit is just the gfp gene whose expression will create fluorescence within the cell. The counter is designed to work like this:
(1) The first arabinose pulse expresses flpe
(2) flpe cuts, flips, and reattaches the whole first SIMM so that now a correctly orientated promoter is leading the next SIMM
(3) This flipping stops flpe transcription and expression because of the gene's inverted orientation with its upstream promoter
(4) Second pulse of arabinose transcribes and expresses cre
(5) Cre cuts, flips, and reattaches the whole second SIMM to place an uninverted promoter upstream of gfp
(6) Cre expression ends just as flpe expression ended earlier
(7) Third arabinose pulse expresses GFP and the cell begins to fluoresce
The two-counter design is identical except without the middle Cre-based step, and thus requiring only two arabinose pulses to express GFP.
The scientists also created another counter by replacing the three arabinose-activated promoters with three different promoters.
Results
The results for these types of counters mirrored those of the RTC counters, those cells received some pulses, but not a full dose had a limited amount of fluorescence, while those receiving a full dosage had a much higher level of fluorescence. This went for both the two and three-counters, as well as being true for both those with only the arabinose-activated promoters and those with the three unique promoters. The explanation for this limited fluorescence is the same as that posed for the RTC counters, that a single pulse expressed the desired protein and a few subsequent proteins.
Figures
Figure 1
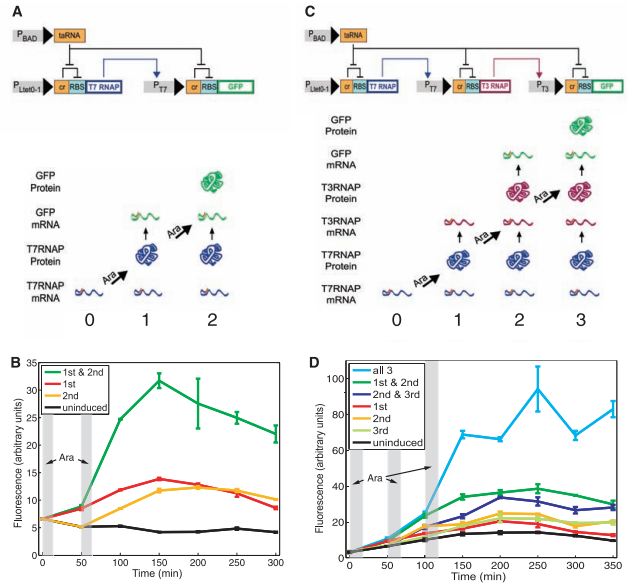
Figure 1A- This is a pictorial representation of the RTC two-counter design on the top and the expected results below that. The design shows the arabinose-activated promoter (P-BAD) upstream of the taRNA, the P-Ltet0-1 promoter-controlled unit with the cis-repressor (cr), ribosome binding site (RBS), and the T7 RNA polymerase gene, and the T7 RNAP controlled unit with a cis-repressor, ribosome binding site, and the GFP gene. The flat headed arrows show the repression interactions occuring, while the normal arrows show the activation interactions. The cr represses the RBS, but the taRNA represses that interaction by binding to the cr, freeing the RBS to be bound to by the ribosome. The gene product for each preceding step transcribes the mRNA for the following unit (the pointed arrow).
The second part of figure 1A pictorial shows what should be the progression to the end point of the counter. The cell starts with T7RNAP mRNAs already transcribed (column 0), so the first arabinose pulse allows the translation of T7 RNAP and its binding to the T7 promoter to transcribe GFP mRNA (column 1). The second arabinose pulse should allow the GFP mRNA to be translated, and make the cell fluoresce.
Figure 1B- This shows the actual experimental results for the RTC two-counter. The two gray-shaded areas represent when the two arabinose pulses occurred. As can be seen from the graph, those cells getting a single pulse showed some fluorescence, but those receiving both had much high fluorescence levels, illustrating that the counters worked.
Figure 1C & D- These serve the same functions as A & B, except for the three-counter instead. The design pictures (C) show the additional step inserted in order to count one higher, but the conventions are the same as A. 1D follows the same conventions as 1B, and the results are simliar in that those cells getting less than a full dose had some fluorescence, but could not match that of the cells receiving 3 pulses.
Figure 2
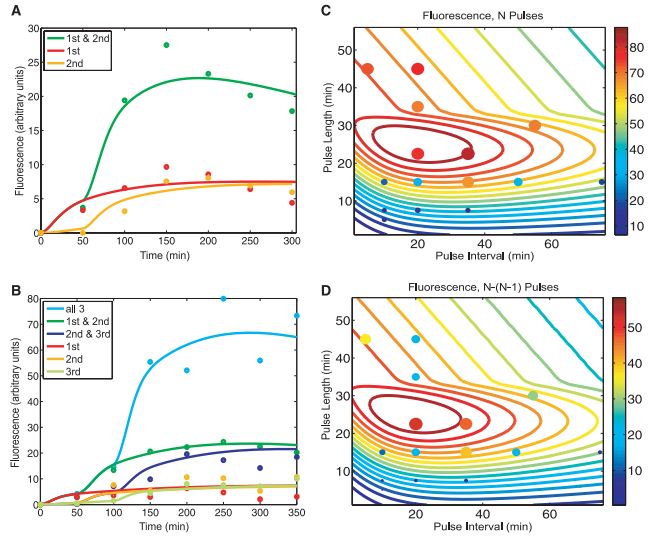
Figure 2A- This is a graph that compares the expected results for the RTC two-counter from the model to the actual experimental results. Each line represents the cells that got that pulse, and the shows the level of fluorescence for that category versus time. The dots are the experimental values measured at specific time points in the actual experiment. The color coding is consistant between the expected and actual lines or dots. The model's predictions closely mirror those of the actual experiment, showing that the model is an accurate one.
Figure 2B- This figure is the same type of figure as 2A, except this one is for the RTC three-counter. Again we see a close match between the predicted values and the experimental ones, as well as the relationships between the different categories being similar.
Figure 2C- The researchers varied the length of the pulses and the time between pulses within their three-counter model, and this figure is a graph of the predicted fluorescence levels as both of those changed. After predicted the results, the authors preformed the actuall experiment and the solid circles are the results of that experiment, following the same color scale as used for the model predictions. These confirmed the model's representation of the counter's ability to function effectively within a relatively large range of values.
Figure 2D- This is the same type of graph as 2C, except what is graphed is the difference in fluorescence between the three-counter and the two-counter at the same conditions. The lines are still predictions for the model, and the solid circles are actual experimental results. This figure further confirmed the ideal range for effective counting using this design.
Figure 3
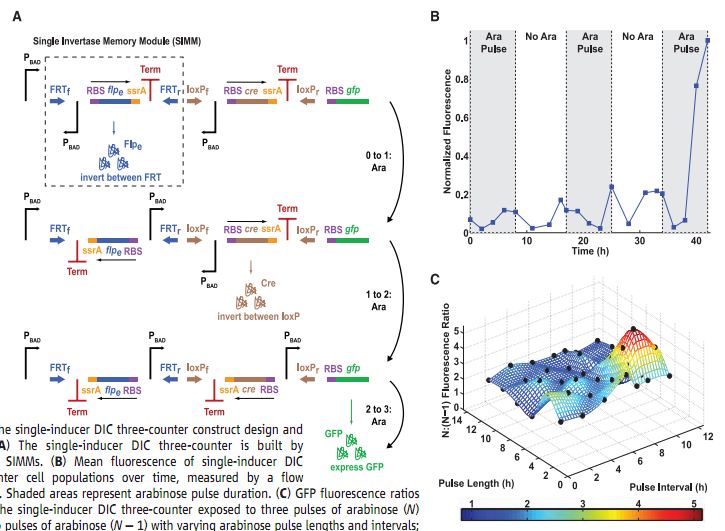
Figure 3A- This is a pictoral representation of the second type of counter, the DIC counter. The individual units that make up the whole counter are the SIMMs, and one is outlined by the dashed box. A SIMM includes an inverted arabinose-activated promoter (P BAD), a RBS, a recombinase gene, a tag for quick degradation of the protein (ssrA), and a transcriptional terminator (Term), all flanked on either side by the cut sites specific for the recombinase encoded within the unit. This specific diagram is for the three-counter and has two SIMMs plus the GFP gene at the end, and uses the flpe, with its FRT cut sites, and the Cre, with its lox cut sites, as the recombinases. The first arabinose pulse transcribes the first SIMM, which is then expressed, cutting and flipping the whole unit at the FRT sites. The second pulse does the same thing for the second SIMM, which cuts at the lox sites instead, but still flips the unit. The third and final pulse transcribes and translates GFP. Each flip correctly orients the P BAD for the transcription of the next unit in the process.
Figure 3B- This is a plot of the experimental results for the DIC three-counter. You can see the small increases in fluorescence in the first 2 pulses due to premature activation of the later process steps, but the increase that comes with the third pulse is by far the largest increase, as it should be if the design functions properly. The shaded regions show the arabinose pulses so that the reader can follow where the counter should be in its process.
Figure 3C- Again the researchers created a mathematical model of their counter design, and used that model to predict the effects of varying pulse length and the time between pulses. What they graphed however, is the ratio of fluorescence levels between the three-counter and the two-counter. The black dots on the graph are experimentally determined results, which closely match the model values. The graph shows ideal ranges of pulse lengths and intervals for the counter to effectively count, which is an important thing to consider when considering uses for the design.
Figure 4
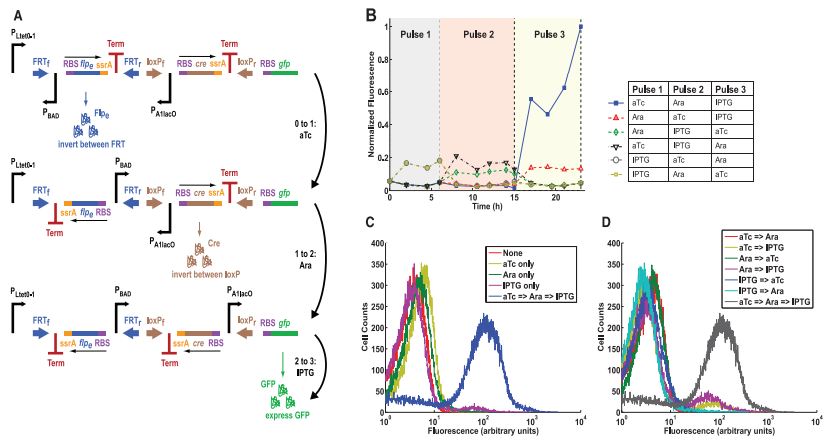
Figure 4A- This is a picture of the DIC counter design that is the same as that shown in Figure 3A, except that this one uses 3 different promoters instead of using P BAD for all of them. In this set-up, the first pulse is anhydrotetracycline (aTc), the second pulse is arabinose, and the third pulse is isopropyl beta-D-1-thiogalactopyranoside (IPTG), each of which activates the promoter for that step, aTc for P Ltet0-1, arabinose for P BAD, and IPTG for P A1lac0.
Figure 4B- This is a graph of flow cytometry data, which measures the level of fluorescence in cells. Every possible combination of the order of the three pulses was tested and the results are shown in this figure. Only those cells that received pulses of aTC, arabinose, and IPTG in that order, had substantial levels of fluorescence at the end of the third pulse, supported the conclusion that the counter design was properly functional.
Figure 4C- This figure is the flow cytometry data comparing those cells that received a single pulse of any of the three activators to those that received all three in the correct order. As can be seen in the graph, only the cells receiving all three pulses had a large number of cells with a high level of fluorescence. The others are all grouped together where cells with low fluorescence fall.
Figure 4D- This figure is similar to 4C, but this time all the cells that received any two pulses are compared to those that received all three in the correct order. Again those cells that got the full dosage have much higher levels of fluorescence than those that only got two of the pulses. This further supports the counter's design functioning as intended.
Opinion
I found this paper and what they were able to make the cells do very interesting, as did friends that I explained it to when asked what I was working on. I wish there had been more explanation for why they performed the N-(N-1) (Figure 2D) analysis, or at least why they included it in the paper over other figures, as I had some trouble understanding what that graph contributed that was not shown in Figure 2C. Especially as the experimental values in Figure 2D only seemed to match the model's predictions in a few places, not as well as they matched for other figures. They also showed the level of fluorescence in multiple different ways between or even within figures. In Figures 1 & 2 the fluorescence is in arbitrary units, but for Figures 3 & 4 it has been normalized to the highest one. The within Figure 4, C & D just show the fluorescence at the end of the process, whereas they've shown how it changed with time for all the cells getting 3 pulses in B. I understand splitting the one, two, and three pulse categories onto different figures so they can be seen better, but why not use the style of B for all three since you can see the final fluorescence level as well as how it changed with time. The paper certainly supports the fact that they created cells that could count to two or three, I just was thrown by the mismatching of some of the figures.
Reference
Friedland AE, Lu TK, Wang X, Shi D, Church G, Collins JJ. Synthetic gene networks that count. Science. 2009; 324:1199-1202.
Synthetic biology uses many tools from other disciplines, or seeks to imitate those tools within biological systems. The researchers in this case wanted to create a synthetic gene network that could be "programmed" to count to two or three like digital circuits can. Possible applications of this ability are being able to program cell death after a certain number of events, or knowing how many times a certain cellular event occurs. The researchers designed and built counters using two different methods: using regulation of the ability of the ribosome to bind to mRNA, and using enzymes that recombine DNA in a known way.
Riboregulated Transcriptional Cascade (RTC) Counters
Method
The counter constructs were created on plasmids that was then inserted into a strain of E. Coli. The two-counters and three-counters were built with the same general method, differing only in how many riboregulated units were involved in the counter set-up, the two-counter had 2 and the three-counter had 3 units. The general unit had, in order, a promoter, a cis-repressor (cr) sequence, a ribosome-binding site (RBS), and a gene. The cr sequence and the RBS were complentary, so when the unit was transcribed, a loop would form as the cr sequence and RBS bound together, blocking the ribosome from landing on the RBS and starting translation. This loop could be undone by a taRNA binding to the cr, freeing the RBS for the ribosome to bind to, starting translation.
Using these facts, the three-counter was designed to work like this:
(1) The cell starts with cr-RBS-T7 RNAP transcripts in it
(2) First pulse of arabinose binds to BAD promoter which transcribes the taRNA
(3) taRNA binds to cr of transcripts, allowing the ribosome to bind to RBS and express T7 RNAP
(4)Pulse ends, arabinose and taRNA are degraded, and T7 RNAP expression ends
(5) Already translated T7 RNAP transcribes cr-RBS-T2 RNAP
(6) Second arabinose pulse creates more taRNA
(7) taRNA binds to cr and frees RBS for ribosome binding and T3 RNAP expression
(8) Pulse ends, arabinose and taRNA are degraded, and T3 RNAP expression ends
(9) T3 RNAP already translated transcribes cr-RBS-GFP
(10) Third pulse of arabinose induces more taRNA
(11) taRNA binds to cr, freeing RBS for ribosome binding, and GFP is expressed
The end result of this design should be a cell that does not fluoresce until after the 3rd arabinose pulse The two-counter uses the same design only without the middle, T3 RNAP step, thus requiring only 2 pulses to express GFP.
Results
The experimental results supported the conclusion that the counter design worked. Cells not given any arabinose did not show any increase in mean fluorescence level. Those given less then a the full amount, whether one pulse for the two-counters or either one or two pulses for the three-counters, showed some increase in fluorescence, but none matched the increase that occurred in cells that received full dosages of arabinose. The limited increase in fluorescence for those cells not getting full doses was explained by the intended protein being expressed, but the cell also expressing a few proteins father down the chain than intended, resulting in fluorescence before it should have occurred.
Model Counters
Methods & Results
Using the design of their RTC counters, the authors built a mathematical model of the counters. They then used these models to make "predictions" for what the experimental results should have been, and analyzed how closely they matched the actual experimental results. The predicted results from the model closely matched those of the real experiments.
Once they authors were confident the model effectively represented what would actually occur, they used it to test the effects of varying the pulse duration and time between pulses. These expected results were also checked against actual results from experiments that varied the pulse duration and gap between them, and again the model closely matched the real-world results. This allowed the researchers to find the maximum and minimum pulse length and frequency, beyond which the counter could not correctly count due to kinetic limits of the cellular processes involved.
DNA Invertase Cascade (DIC) Counters
Method
The DIC counters were built using well-known and understood DNA recombinases Cre and flpe built into units called single invertase memory modules (SIMMs). Stringing together different numbers of these SIMMs on a plasmid allow the cells to count to either two or three depending on how many SIMMs the plasmid includes. The design of the three-counter is 2 SIMMs followed by the GFP gene. The 1st SIMM is the gene for flpe recombinase led by an inverted arabinose-activated promoter, and flanked by sites where flpe can cut, invert the DNA segment, and reattach in the new orientation. The 2nd SIMM is similar except being the cre gene and being flanked by the restriction sites for Cre, but with the same inverted arabinose-activated promoter. The last unit is just the gfp gene whose expression will create fluorescence within the cell. The counter is designed to work like this:
(1) The first arabinose pulse expresses flpe
(2) flpe cuts, flips, and reattaches the whole first SIMM so that now a correctly orientated promoter is leading the next SIMM
(3) This flipping stops flpe transcription and expression because of the gene's inverted orientation with its upstream promoter
(4) Second pulse of arabinose transcribes and expresses cre
(5) Cre cuts, flips, and reattaches the whole second SIMM to place an uninverted promoter upstream of gfp
(6) Cre expression ends just as flpe expression ended earlier
(7) Third arabinose pulse expresses GFP and the cell begins to fluoresce
The two-counter design is identical except without the middle Cre-based step, and thus requiring only two arabinose pulses to express GFP.
The scientists also created another counter by replacing the three arabinose-activated promoters with three different promoters.
Results
The results for these types of counters mirrored those of the RTC counters, those cells received some pulses, but not a full dose had a limited amount of fluorescence, while those receiving a full dosage had a much higher level of fluorescence. This went for both the two and three-counters, as well as being true for both those with only the arabinose-activated promoters and those with the three unique promoters. The explanation for this limited fluorescence is the same as that posed for the RTC counters, that a single pulse expressed the desired protein and a few subsequent proteins.
Figures
Figure 1
Figure 1A- This is a pictorial representation of the RTC two-counter design on the top and the expected results below that. The design shows the arabinose-activated promoter (P-BAD) upstream of the taRNA, the P-Ltet0-1 promoter-controlled unit with the cis-repressor (cr), ribosome binding site (RBS), and the T7 RNA polymerase gene, and the T7 RNAP controlled unit with a cis-repressor, ribosome binding site, and the GFP gene. The flat headed arrows show the repression interactions occuring, while the normal arrows show the activation interactions. The cr represses the RBS, but the taRNA represses that interaction by binding to the cr, freeing the RBS to be bound to by the ribosome. The gene product for each preceding step transcribes the mRNA for the following unit (the pointed arrow).
The second part of figure 1A pictorial shows what should be the progression to the end point of the counter. The cell starts with T7RNAP mRNAs already transcribed (column 0), so the first arabinose pulse allows the translation of T7 RNAP and its binding to the T7 promoter to transcribe GFP mRNA (column 1). The second arabinose pulse should allow the GFP mRNA to be translated, and make the cell fluoresce.
Figure 1B- This shows the actual experimental results for the RTC two-counter. The two gray-shaded areas represent when the two arabinose pulses occurred. As can be seen from the graph, those cells getting a single pulse showed some fluorescence, but those receiving both had much high fluorescence levels, illustrating that the counters worked.
Figure 1C & D- These serve the same functions as A & B, except for the three-counter instead. The design pictures (C) show the additional step inserted in order to count one higher, but the conventions are the same as A. 1D follows the same conventions as 1B, and the results are simliar in that those cells getting less than a full dose had some fluorescence, but could not match that of the cells receiving 3 pulses.
Figure 2
Figure 2A- This is a graph that compares the expected results for the RTC two-counter from the model to the actual experimental results. Each line represents the cells that got that pulse, and the shows the level of fluorescence for that category versus time. The dots are the experimental values measured at specific time points in the actual experiment. The color coding is consistant between the expected and actual lines or dots. The model's predictions closely mirror those of the actual experiment, showing that the model is an accurate one.
Figure 2B- This figure is the same type of figure as 2A, except this one is for the RTC three-counter. Again we see a close match between the predicted values and the experimental ones, as well as the relationships between the different categories being similar.
Figure 2C- The researchers varied the length of the pulses and the time between pulses within their three-counter model, and this figure is a graph of the predicted fluorescence levels as both of those changed. After predicted the results, the authors preformed the actuall experiment and the solid circles are the results of that experiment, following the same color scale as used for the model predictions. These confirmed the model's representation of the counter's ability to function effectively within a relatively large range of values.
Figure 2D- This is the same type of graph as 2C, except what is graphed is the difference in fluorescence between the three-counter and the two-counter at the same conditions. The lines are still predictions for the model, and the solid circles are actual experimental results. This figure further confirmed the ideal range for effective counting using this design.
Figure 3
Figure 3A- This is a pictoral representation of the second type of counter, the DIC counter. The individual units that make up the whole counter are the SIMMs, and one is outlined by the dashed box. A SIMM includes an inverted arabinose-activated promoter (P BAD), a RBS, a recombinase gene, a tag for quick degradation of the protein (ssrA), and a transcriptional terminator (Term), all flanked on either side by the cut sites specific for the recombinase encoded within the unit. This specific diagram is for the three-counter and has two SIMMs plus the GFP gene at the end, and uses the flpe, with its FRT cut sites, and the Cre, with its lox cut sites, as the recombinases. The first arabinose pulse transcribes the first SIMM, which is then expressed, cutting and flipping the whole unit at the FRT sites. The second pulse does the same thing for the second SIMM, which cuts at the lox sites instead, but still flips the unit. The third and final pulse transcribes and translates GFP. Each flip correctly orients the P BAD for the transcription of the next unit in the process.
Figure 3B- This is a plot of the experimental results for the DIC three-counter. You can see the small increases in fluorescence in the first 2 pulses due to premature activation of the later process steps, but the increase that comes with the third pulse is by far the largest increase, as it should be if the design functions properly. The shaded regions show the arabinose pulses so that the reader can follow where the counter should be in its process.
Figure 3C- Again the researchers created a mathematical model of their counter design, and used that model to predict the effects of varying pulse length and the time between pulses. What they graphed however, is the ratio of fluorescence levels between the three-counter and the two-counter. The black dots on the graph are experimentally determined results, which closely match the model values. The graph shows ideal ranges of pulse lengths and intervals for the counter to effectively count, which is an important thing to consider when considering uses for the design.
Figure 4
Figure 4A- This is a picture of the DIC counter design that is the same as that shown in Figure 3A, except that this one uses 3 different promoters instead of using P BAD for all of them. In this set-up, the first pulse is anhydrotetracycline (aTc), the second pulse is arabinose, and the third pulse is isopropyl beta-D-1-thiogalactopyranoside (IPTG), each of which activates the promoter for that step, aTc for P Ltet0-1, arabinose for P BAD, and IPTG for P A1lac0.
Figure 4B- This is a graph of flow cytometry data, which measures the level of fluorescence in cells. Every possible combination of the order of the three pulses was tested and the results are shown in this figure. Only those cells that received pulses of aTC, arabinose, and IPTG in that order, had substantial levels of fluorescence at the end of the third pulse, supported the conclusion that the counter design was properly functional.
Figure 4C- This figure is the flow cytometry data comparing those cells that received a single pulse of any of the three activators to those that received all three in the correct order. As can be seen in the graph, only the cells receiving all three pulses had a large number of cells with a high level of fluorescence. The others are all grouped together where cells with low fluorescence fall.
Figure 4D- This figure is similar to 4C, but this time all the cells that received any two pulses are compared to those that received all three in the correct order. Again those cells that got the full dosage have much higher levels of fluorescence than those that only got two of the pulses. This further supports the counter's design functioning as intended.
Opinion
I found this paper and what they were able to make the cells do very interesting, as did friends that I explained it to when asked what I was working on. I wish there had been more explanation for why they performed the N-(N-1) (Figure 2D) analysis, or at least why they included it in the paper over other figures, as I had some trouble understanding what that graph contributed that was not shown in Figure 2C. Especially as the experimental values in Figure 2D only seemed to match the model's predictions in a few places, not as well as they matched for other figures. They also showed the level of fluorescence in multiple different ways between or even within figures. In Figures 1 & 2 the fluorescence is in arbitrary units, but for Figures 3 & 4 it has been normalized to the highest one. The within Figure 4, C & D just show the fluorescence at the end of the process, whereas they've shown how it changed with time for all the cells getting 3 pulses in B. I understand splitting the one, two, and three pulse categories onto different figures so they can be seen better, but why not use the style of B for all three since you can see the final fluorescence level as well as how it changed with time. The paper certainly supports the fact that they created cells that could count to two or three, I just was thrown by the mismatching of some of the figures.
Reference
Friedland AE, Lu TK, Wang X, Shi D, Church G, Collins JJ. Synthetic gene networks that count. Science. 2009; 324:1199-1202.