This web page was produced as an assignment for an undergraduate course at Davidson College.
"Coat Variation in the Domestic Dog is Governed by Variants in Three Genes"
Cadieu et al.
Summary
The aim of this research was to find any genetic correlation among variation in dog coat length, curl, and pattern. The researchers selected one data set (the dachshund data set) composed of one breed with three observably different coat phenotypes in order to reduce variables (as in one of the few genetic differences would result in different coat phenotype), chose another data set that was composed of breed with distinctly curly or not curly hair (the Portuguese Water Dog (PWD) data set), again to reduce variation, and compared these two data sets against a data set comprising a large number of breeds encompassing a large number of phenotypes to ensure that said variation occurred across many phenotypes, named the CanMap data set.
In order to compare these data sets the researchers conducted a genome wide association study (GWAS), which is used to determine the association between single nucleotide polymorphisms (SNPs) at a specific locus with a certain phenotype. If a significantly strong correlation between a SNP (representing a certain mutation) and a phenotype was observed, one could say that the observed phenotype was due to having that mutation.
The researchers in fact found significant correlations between mutations in the RSPO2, FGF5, and KRT71 genes with having furnishings (eyebrows and mustache), long hair, and curly hair, respectively. The evidence for these correlations and what exactly the researchers found will be discussed below in the figure analysis portion.
The ultimate implication of this research is that seemingly complex and intricate phenotypes can be reduced to mutations in a few genes. The authors also speculated that these few mutations arose in few individuals, and rapidly appeared in many breeds, speaking to the power of artificial selection. They claim that, possibly, more differences in phenotypes can be explained by few genes.
Figure Analysis
Figure 1
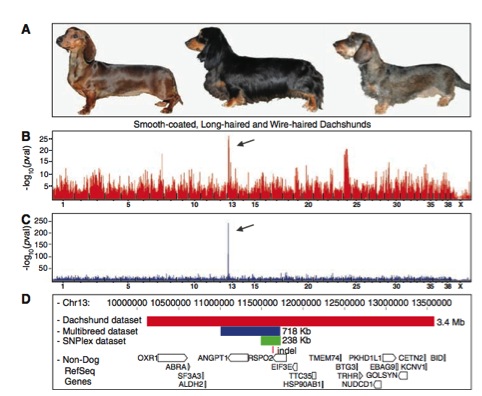
This figure portrays the correlation between a mutation on RSPO2 and having furnishings. A This box pictorially shows the three phenotypes of the kinds of dogs the composed the first data set. B This box shows the GWAS analysis for wire-haired dachshunds compared to the smooth-coated and long-haired dachshunds (which served as controls). Plotted on the y-axis is the most significant (p-value through a chi-squared test) LOD score (the higher this value, the more likely the loci is linked to the phenotype) and on the x-axis is locus position (numbers indicate chromosome). The arrow points to th locus that is most likely correlated with having furnishings. C This box similarly portrays data as in panel B, but compares the dachshund data set against the CanMap data set. Once again a strong correlation arises with a SNP on chromosome 13. This panel eliminates many false correlations that arose due to similarities within the same breed. Box D displays the results of the fine mapping of chromosome 13 amongst the data set. The red box displays the common haplotype (group of alleles often inherited together) for dachshunds with furnishings, which correlates to the loci indicated in Panels A and B. The blue rectangle represents the haplotype associated with having furnishings in the CanMap dataset. Notice it is smaller than in the dachshund data set, because more recombination occurs amongst various breeds, thus the haplotype shrinks. The green box displays the furnishing associated haplotype of yet another data set, further homing in on the specific allele associated with having furnishings. Data shown on another table (not shown in this report, due to special constraints) indicated the culprit allele contained as a 167 base pair insertion on the 3’ untranslating region of the RSPO2 gene displayed in panel D (the insertion in vertically above genes found in this region of chromosome 13). This insertion causes an overproduction of the protein, resulting in furnishings.
Figure 2
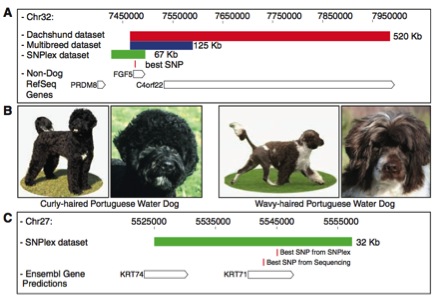

This figure shows the correlation between having a mutation on the FGF5 and KRT71 genes, and having long and curly hair, respectively. Panel A, similarly to figure 1, shows haplotypes of datasets associated with having long hair. The most likely SNP associated with having long hair, found with GWAS, analysis is shown as lying on the FGF5 gene (because the graph is vertically synchronized). Box B pictorially displays the curly- and wavy-haired portugese water dogs. Panel C shows the haplotype associated with having curly hair (the green rectangle) and two most likely SNPs obtained from conducting GWAS analysis (graphs not shown) using two different control populations. Both SNPs are shown to lie on gene KRT71, indicating a mutation in this gene results in curly hair.
Figure 3
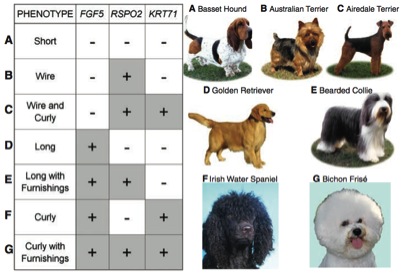
This figure depicts the possible combinations of mutations (+) and ancestral strains (-) a dog could have in the three genes discussed in the paper. The pictures (right) pictorially represent the dogs with the genotype shown at the right. For example, the basset hound, labeled “A” on both sides of the graph has ancestral (-) genes at all three genes, resulting in a short hair phenotype. The implication of this figure is that the combination of mutations in few (three) genes can result in the seemingly much different phenotypes (pictured right).
Conclusions
This paper demonstrated that variation in alleles for a few genes can result in a plethora of phenotypes. It also taught me about various statistical analyses for associating SNPs with phynotypes, which can obviously have very useful applications. This paper may inspire others to attempt to break down a seemingly complex phenotype into investigating a few genes, by breaking down the phenotype one component at a type.
References
All images from http://bio.davidson.edu/courses/genomics/2011/Bio309_papers/Dog_coats.pdf
http://ghr.nlm.nih.gov/handbook/genomicresearch/gwastudies# GWAS information
http://ghr.nlm.nih.gov/glossary=singlenucleotidepolymorphism SNP information
http://en.wikipedia.org/wiki/Genetic_linkage#LOD_score_method_for_estimating_recombination_frequency LOD score information
Davidson Genomics
Home
Questions? Comments? Contact me at mike.nuttle@comcast.net


