This web page was produced as an assignment for an undergraduate course at Davidson College.
Researchers identify genes potentially responsible for differences in wild and domestic chicken

Image courtesy of animalEquality.
1) What was the research project?
A 2012 study published in Genomics compared the transcriptomes of two breeds of domesticated chicken (chahua chicken and avian broiler) and one breed of wild chicken (red jungle fowl) using 2 genomics technologies to gain insight into the genes that are being differentially expressed between species (Li, et al., 2012). This would allow us to better select (or engineer) livestock with economically favorable phenotype at a molecular basis. They found 7 genes that were differentially expressed in the domestic breeds and based on their associated functions the group determined they had found a pool of candidate genes.
2) Were they testing a hypothesis or doing discovery science?
This project was discovery science, because they did not go into the project with a hypothesis they were testing. The group’s goal, as they stated, was to contribute to the “comprehensive understanding of the molecular mechanism of animal domestication” (Li, et al., 2012). They were simply searching for differences that they could characterize in the genomes of wild versus domestic chickens.
3) What genomic technology was used in the project?
The transcriptomes of each species were analyzed using RNASeq data from Illumina HiSeq 2000 (Li, et al. 2012). After the initial analyses of the data, 11 genes were randomly selected and subjected to real time PCR validation. They compared the data using the Spearman's rank correlation coefficient and found that the RT PCR validated their RNASeq analysis data.
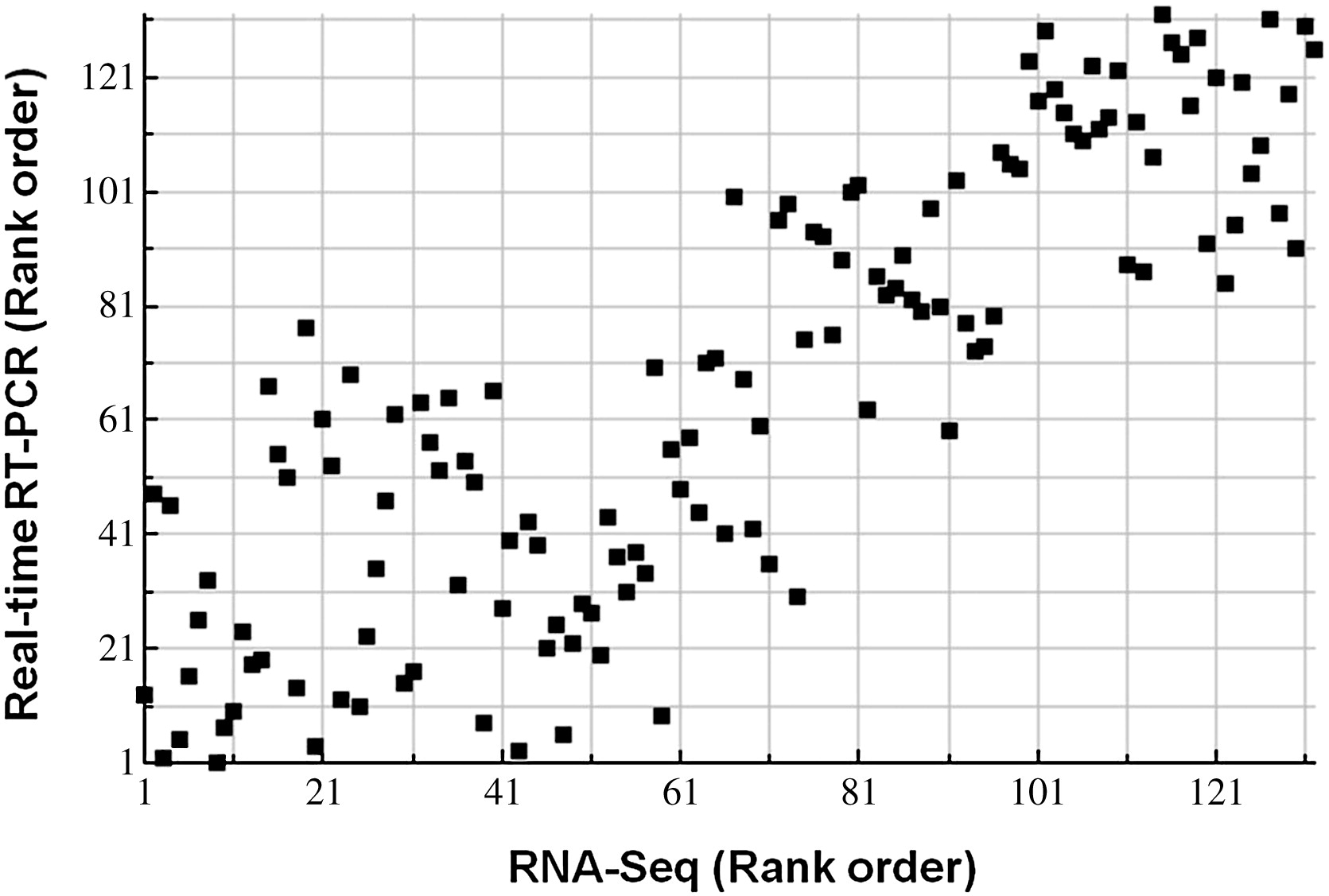
Fig. 1.
Scatter plot of correlation analysis between the RNA-seq data and real-time RT
PCR data. The consistency between RNA-seq and real-time RT PCR results were determined
by calculating the Spearman's rank correlation coefficient for the expression of 11 randomly
selected genes across 12 samples (4 samples for each breeds, total n = 132). (replicated Li, et al., 2012)
4) What was the take home message?
The study found 297 genes that were up regulated and 262 genes that were down regulated in both chahua and avian broiler as compared to the red jungle fowl (Li, et al. 2012). The 7 commonly differentially expressed genes the study focused on, whether they were up-regulated or down-regulated, were biologically important and related to known function and easily connected to the favored phenotypes. These genes expression differences displayed are evidently significant and related to muscle development and growth regulation in chickens.
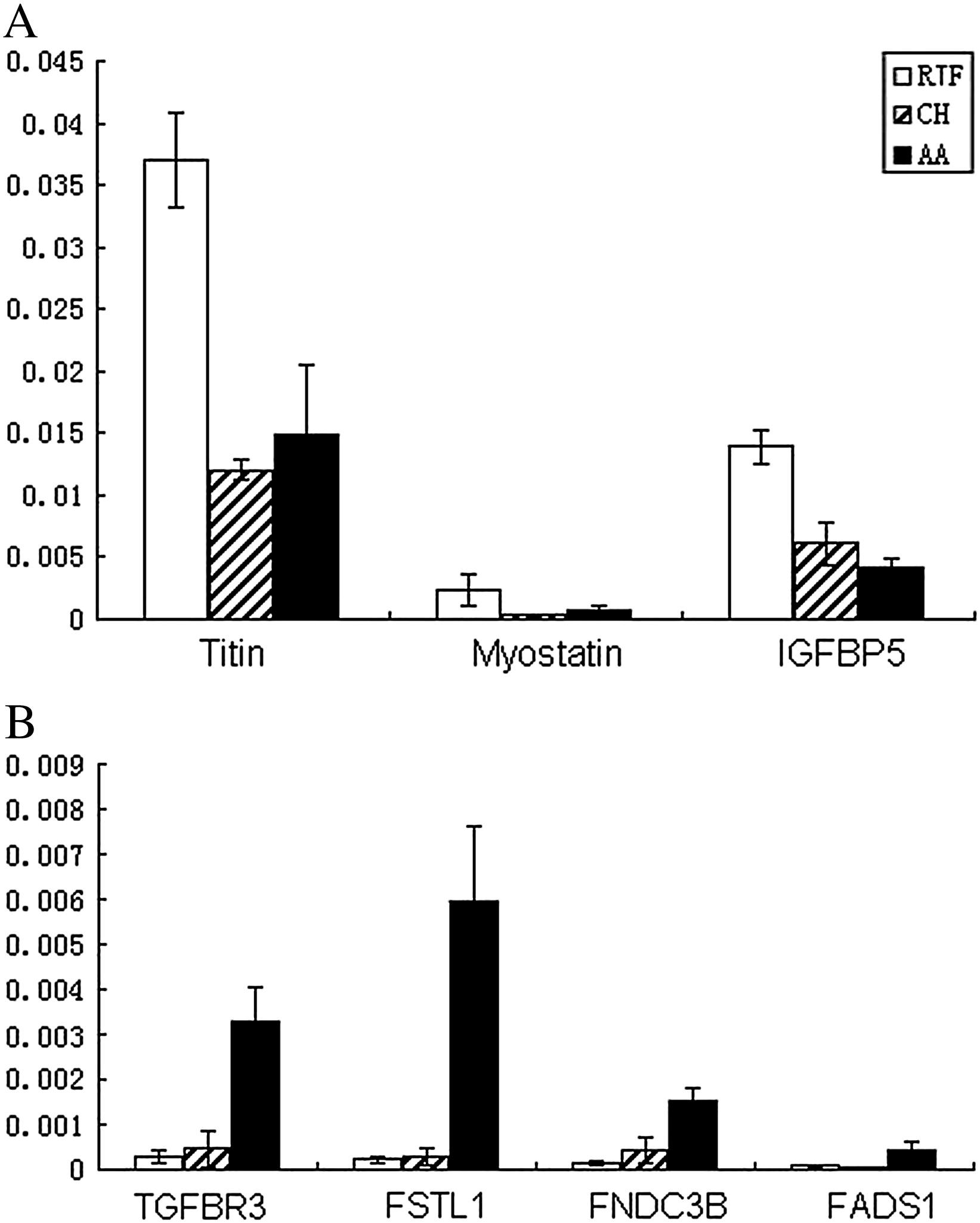
Fig. 3.
Real-time RT PCR validation of biologically important genes that differentially
expressed between the wild and domestic chicken breeds. Gastrocnemius from 5
chickens was used in this experiment, and expression levels of genes were normalized
to that of GAPDH. Q-PCR results were analyzed with 2△△Ct method. A: expression of
down-regulated genes in domestic breeds compared with the RJF. B: expression of
up-regulated genes in domestic breeds. (replicated Li, et al., 2012)
The study established an expression difference basis for the phenotypic differences that we can see in wild versus domestic chickens. It also demonstrated that selection of these genes that drive increased muscle mass and rapid growth may also be driving selection for other intracellular processes that accompany these processes. In this case it is specifically the ubiquitin proteasome pathway.
These genes and information about them could are important and can be used to inform and potentially enhance breeding methods. Also, the study demonstrated the effect that human selective pressures could have potentially had on the expression of genes in chickens, which can be informative in what factors are affecting selection over time.
5) What is your evaluation of the project?
In recent years, there has been much controversy surrounded genetically modified organisms (GMOs) and whether to introduce genetically modified livestock into the market. Determining the exact genes responsible for increased muscle mass would be of great interest to livestock companies. It could be easier to convince the public the safety of further enhancing genes that have been previously selected for over centuries within a species rather than introduction of a different species DNA and crossing species (Marris 2001). The development of these economically favorable phenotypes would also provide more meat production and increased profit for livestock companies, making this research of great economic interest.
The differentially expressed genes discovered could also be of value in clinical and other general research by knowing genes that are responsible for these desired phenotypes. We may be able to find the orthologous gene in humans and further study these. These genes could be a useful therapy treatment to combat the effects of muscular dystrophy and other muscular degenerative disease.
In all, I believe the work is important and the research should be continued. I think that verification of this experiment would be valuable and that the groups methodology was sometimes confusing, but the effectively conveyed their message.
Scientific Evaluation:
The researchers mention that they conducted some breed comparisons with other chicken breeds, but the breeds are not stated. The species Gallus gallus has 6 subspecies 2 of which are studied here. In the article it is never specifically stated which subspecies group is being used. It seems reasonable to believe the 2 domestic breeds belong to Gallus gallus domesticus, but there are also 5 other subspecies groups native to different parts of Asia. The research group is based in China, so while not explicitly stated, it also seems reasonable to believe that the red jungle fowls used belong to Gallus gallus gallus. Noting this would’ve been helpful to a reader or researcher that was interested in recreating this experiment. Even if these assumptions are incorrect, it would be beneficial to collect members of each of the subspecies and compare the expression profiles within the species. We know that environmental pressures are a driving force in selection, and the defining characteristic of a subspecies is geographic location. There could be another subspecies population that has a more similar expression profile to the 2 domestic breeds, which might be suggestive that the domestic populations are more related to a particular subspecies than to the others. This could suggest that the differential gene expression is not due to human selective pressures, or shed more light on the mechanisms of selection. Even if the domestic breeds were not more similar to some other subspecies, it would just serve as a control and reinforce the group’s findings. I think this would be a beneficial addition to the research.
The group obtained preliminary results and then used a secondary method for validation of their data. However, the rational behind their sampling methods is not clear to the reader in some instances. It is hard for the reader to validate the significance of some of their sampling techniques when the rationale is not outlined. It is also difficult to determine how the leap was made from this large number of genes that were differentially expressed to 7 genes and they provide no reasoning for how the search was narrowed. It is likely that they selected the genes that seemed directly connected to the displayed phenotypes based on the GO enrichment analysis, but all the genes are not provided, so verification cannot take place. I do think that further studies done with various other tissues, as all the samples were only taken from the gastrocnemius would be valuable. Transcriptomics explores the RNA contained in a given cell or tissue. For this reason, it would be beneficial to see if the same genes are differentially expressed in the pectoral muscles, which humans have also heavily selected. If the two tissues show the same genes have been differentially expressed, it would be a good verification of these findings.
References
Marris, C. 2001. Public views on GMOs: deconstructing the myths. EMBO Rep. [Internet]. [Cited 30 Jan 2016] 2 (7): 545-548. Available at:http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1083956/
Email Questions or Comments: duatchley@davidson.edu
Genomics Page Dustin's Home Page © Copyright 2016 Department of Biology, Davidson College,
Davidson, NC 28035