Figure 1. A high magnification
of yeast cells reproducing by budding
(<http://www.barrelhouse.com/yeast.html>).
Figure 1. A high magnification
of yeast cells reproducing by budding
(<http://www.barrelhouse.com/yeast.html>).
Both my favorite annotated yeast
gene, CDC28, and my favorite non-annotated yeast gene, YBR161W are
located on Chromosome II of Saccharomyces cerevisiae:
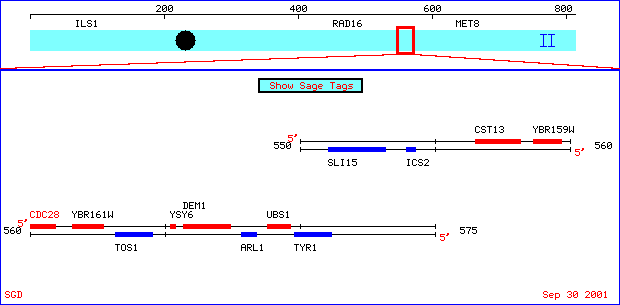
Figure 2. Close-up of chromosome
II including my two favorite yeast genes, CDC28 and YBR161W. With
permission of SGD <http://genome-www4.stanford.edu/cgi-bin/SGD/ORFMAP/ORFmap?sgdid=S0000364>.
My Favorite Annotated Yeast Gene: CDC28
A Little Background Information on CDC28:
One of biology's fundamental truisms is that all cells come from pre-existing cells. As such, every cell has a highly evolved and complex signaling pathway to undergo mitosis, the cell division process that produces identical nuclei for all the cells of an adult organism's body. There are two phases in the life of a cell: mitosis and interphase. Interphase consists of three subphases: G1, immediately following mitosis, S, DNA replication, and G2, another inactive state that precedes mitotic division. The protein that controls a cell's "decision" to replicate its DNA and enter mitosis is called a cyclin-dependent kinase (Cdk). It is the generic enzyme that catalyzes the phosphorylation of certain amino acids in different proteins, playing an essential role in initiating each phase of the cell cycle. Cdk is not active by itself; it must be bound by a second type of proteins called cyclins, hence the cyclin dependence. The activation and deactivation of this complex system make up the finely tuned machinery which is responsible for cell division and growth (Purves et al., 1998).
So where does CDC28 come in?
In Saccharomyces cerevisiae, CDC28, or Cdk1, is the mother of all enzymes in mitotic division: it is the single Cdk that interacts with two families of cyclins throughout the major cell cycle events of a budding yeast. Cdk activity is controlled by the alternating creation and degredation of the 9 cyclins (Cln 1-3 and Clb 1-6) that CDC28 catalyzes (Chen et al., 2000; <http://www.molbiolcell.org/cgi/content/full/11/1/369?view=full&pmid=10637314>). The stage-specific activators are Cln1-3 (G1 phase), Clb 5,6 (S phase), Clb 3,4 (S/G2 phase) and Clb 1,2 (mitosis) (Kuntzel et al., 1996; <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=8922282&dopt=Abstract>).
How well conserved is CDC28 across
species?
CDC28
and the complex signaling pathway associated with it is highly conserved
across species. Comparing yeast's CDC28 to mammalian homologs yielded
similar proteins and pathways in humans, mice and rats with very high E-values
(SGD, 2001; <http://genome-www.stanford.edu/cgi-bin/SGD/Sacch3D/getblast?name=CDC28&db=mammal>).
A protein Entrez search for CDC28 yielded similar proteins in Caenorhabditis
elegans, Branchiostoma belcheri,
Arabidopsis thaliana, Mus musculus, and Homo sapiens (Entrez,
2001; <http://www.ncbi.nlm.nih.gov:80/entrez/query.fcgi?CMD=search&DB=protein>).
Once an effective system has evolved, slight variations of it are typically is seen in many different
types of organisms.
What happens if yeast can't make a
functional copy of CDC28?
A deletion of CDC28 is a serious problem; an extensive and systematic deletion study found that without it, yeast cells were inviable. Without CDC28, cells cannot replicate and grow, and life ends quickly following its beginning (Winzeler et al., 1999; <http://genome-www4.stanford.edu/cgi-bin/SGD/phenotype/phenotype.pl>).
So why is it important to understand
CDC28 and how it works?
Not only is
cell division and growth an essential physiological process that affects
all organisms, this pathway is of particular interest to cancer researchers.
If we can better understand how normal tissue grows and divides, we can
better contrast it to abnormal and out of control cell division characteristic
of cancerous tumors.
My Favorite Non-Annotated Yeast Gene: YBR161W
YBR161W is an Open Reading Frame (ORF) located on Chromosome II of Saccharomyces cerevisiae, in close proximity to the CDC28 gene. Both the nucleotide and amino acid sequences for this ORF can be seen here (Entrez, 2001; <http://www.ncbi.nlm.nih.gov:80/entrez/query.fcgi?cmd=Retrieve&db=nucleotide&list
BLAST nucleotide, Conserved Domain and Protein Data Bank searches all yielded no new information.
A protein BLAST search
of the amino acid sequence for YBR161W yielded several hits for the sur1
protein in fission yeast (<http://www.ncbi.nlm.nih.gov/blast/Blast.cgi>).
The hits with the best identity and positive
percentages, as well as the highest E-values, indicated YBR161W may be a possible functional
homolog to the transmembrane protein coded for by the Sur1 gene on chromosome
XVI.
What is Sur1?
Sur1, according
to a recent study, is a gene required for Saccharomyces
cerevisiae growth in a calcium-rich medium; Sur1 mannosylates inositolphosphorylceramide,
a component of cell wall formation (Beeler et al., 1997;<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids=9323360&dopt
=Abstract>).
Searching SGD for Sur1 revealed that this protein is involved in maintenance
of phospholipid levels through mannose-inositol-P-ceramide (MIPC) metabolism
(SGD, 2001; <http://genome
-www4.stanford.edu/cgi-bin/SGD/locus.pl?locus=sur1>).
Phospholipids make up the central component of cell walls; thus, Sur1 plays
an important role in the generation of new cells.
Kyte-Doolittle Analysis:
A hydropathy plot analysis of YBR161W revealed two peaks at more than 2.5, confirming the potential for a transmembrane domain (<http://fasta.bioch.virginia.edu/fasta/cgi/pgrease.cgi>).
Conclusions:
Based on sequence
similarity, the ORF YBR161W may be a functional homolog to the yeast gene
Sur1, which has been found to play a key role in cell wall formation.
An important piece of information is missing, however: whether or not the
ORF itself actually codes for protein. PCR and Western Blot
analysis could better answer this question. If the ORF does code
for protein, more experimentation could determine if it also plays a role
in cell wall formation and subsequent cellular fission.
An interesting consequence
of choosing an annotated gene and an ORF right next to each other on the
chromosome has been to see if they play similar roles in known cellular processes. It appears
that my ORF has a similar function to my gene; they may both be
involved in the cellular processes of growth and proliferation.
1. Beeler TJ, Fu D, Rivera
J, Monaghan E, Gable K, Dunn TM. 1997. SUR1 (CSG1/BCL21), a
gene necessary for growth of Saccharomyces cerevisiae in the presence of
high Ca2+
concentrations at 37 degrees C, is required for mannosylation of inositolphosphorylceramide.
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&list_uids
=9323360&dopt=Abstract>. Accessed 2001 Sept 28.
2. Chen KC, Csikasz-Nagy
A, Gyorffy B, Val J, Novak B, TYson JJ. 2000. Kinetic analysis of a
molecular model of the budding yeast cell cycle: a consensus picture of
cell cycle controls
in budding yeast. <http://www.molbiolcell.org/cgi/content/full/11/1/369?view=
full&pmid=10637314>. Accessed 2001 28 Sept.
3. Kuntzel H, Schulz A,
Ehbrecht IM. 1996. Cell cycle control and initiation of DNA replication
in Saccharomyces cerevisiae. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd
=Retrieve&db=PubMed&list_uids=8922282&dopt=Abstract>).
Accessed 2001 Sept 28.
4. Purves WK, Orians GH,
Heller HC, Sadava D. 1998. Life: The Science of Biology. Sinauer
Associates, Inc. p 196-197.
5. SGD. BLASTPed against
mammalian protein sequences in GenBank (11 Aug 1997).
<http://genome-www.stanford.edu/cgi-bin/SGD/Sacch3D/getblast?name=CDC28&db
=mammal>. Accessed 2001 Sept 28.
6. Winzeler EA, Shoemaker
DD, Astromoff A, Liang H, Anderson K, Andre B, Bangham R,
Benito R, Boeke JD, Bussey H, Chu AM, Connelly C, Davis K, Dietrich F,
Dow SW, El
Bakkoury M, Foury F, Friend SH, Gentalen E, Giaever G, Hegemann JH, Jones
T, Laub M,
Liao H, Davis RW, et al. 1999. Functional characterization of the S. cerevisiae
genome by
gene deletion and parallel analysis.
<http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db
=PubMed&list_uids=10436161&dopt=Abstract>. Accessed 2001
Sept 28.
Department of Biology, Davidson College, Davidson, NC 28036
Send
comments, questions, and suggestions to: emoldham@davidson.edu