Here is a series of step-by-step figures that can help you remember how chromosomal walking is performed.
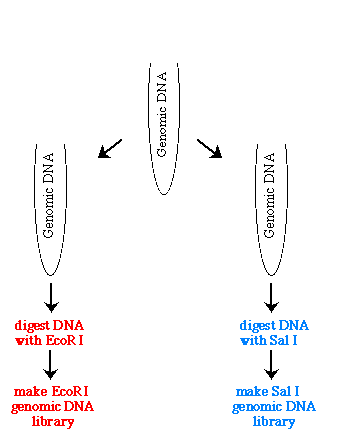
The first thing you need to do is create two genomic libraries of the same DNA but each library used a different restriction enzyme, such as EcoR I and Sal I.

Figure 1. Illustration of how one genome is used to prepare two genomic libraries with two different restriction enzymes.
Next, you must have a probe that is linked to your favorite gene (YFG). In this example, we want to clone the cystic fibrosis gene (CF). You would screen a genomic library with this probe and isolate a piece of DNA that binds to your probe. If you cloned an EcoR I restriction fragment from the genomic library (let's say it is fragment #3 of many possible fragments) that binds to a particular probe (called MET) that is linked to CF. You want to slide down the chromosme from MET towards CF so you can clone and sequence CF. D7S8 is another RFLP marker located on the other side of CF so CF is located between MET and D7S8.
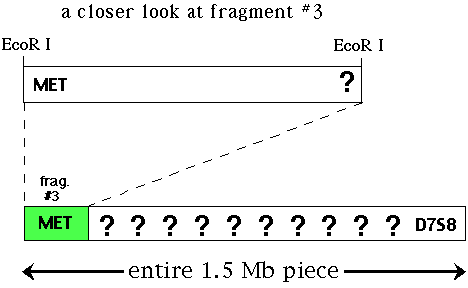
Figure 2. Illustration of one EcoR I fragment that binds to the probe MET. Located at the other end is a second RFLP marker called D7S8 and CF is located somewhere between these two markers.
To clone CF, you will employ chromosomal walking to take baby steps towards CF, starting with the EcoR I restriction fragment #3 you just cloned. Now, you need to generate a restriction map of EcoR I fragment #3. You must digest the Eco RI restriction fragment with multiple restriction enzymes and analyse the results on an agarose gel as shown here:
Figure 3. Cartoon of an agarose gel that contains the restricted DNA as described above.
If you then performed a Southern blot with this gel and used the original MET probe that allowed you to isolate the EcoR I restricion fragment, you might see the following resutls on an X-ray film:
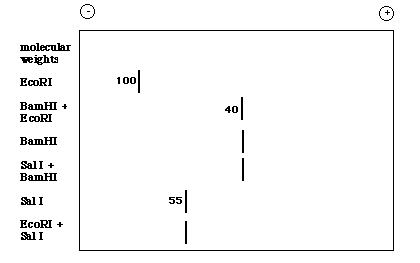
Figure 4. Cartoon of the X-ray film obtained when the Southern blot is probed with MET.
From these data, you could construct the following restriction map that also indicates where the probe binds (note: two slightly different restriction maps could be generated from these data but this one is fine for our purposes):
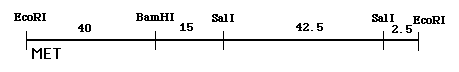
Figure 5. One possible restriction map, given the data from figures 1 and 2 above.
The next task at hand is to isolate the 2.5 kb Sal I - EcoR I fragment and use it as your second probe on the Sal I genomic library because this 2.5 kb piece is the DNA fragment furthest from the MET marker and therefore must be closer to CF. You are ready to screen the Sal I genomic library that used identical DNA but was digested with the restriction enzyme Sal I instead of EcoR I. Because probe #2 is flanked on the left by a Sal I site, you know any new fragment that has Sal sites on both ends and binds to the second probe will extend towards the right (in the direction of CF) as shown:
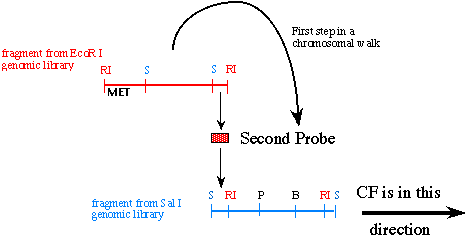
Figure 6. This cartoon illustrates how you use the first genomic DNA clone to generate a second probe that takes one step in the direction of CF.
When you have cloned a Sal I fragment that binds to probe #2, you need to figure out its restriction map the same way we did for the EcoRI fragment #3 above. This process continues until you reach D7S8. The final product pf a chromosomal walk is a series of overlapping restriction maps starting at your original probe (MET) and extending to D7S8. The final combined restriction map, and the overlapping fragments, might look like this:
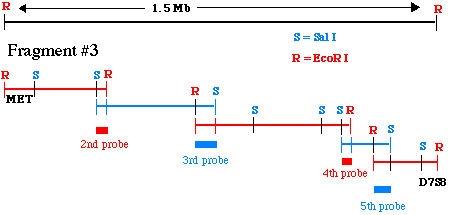
Figure 7. This cartoon illustrates how
a series of overlapping pieces of genomic DNA has been isolated
from alternate genomic libraries.
This is a simplistic example involving only 5 steps for the walk.
Now that you have a restriction map spanning the area of interest, you can use a number of different methods to determine which fragment contains the CF gene. Once you know this, it is very simple to sequence the CF gene and continue your analysis.
Although this example used CF, chromosomal walking can be used to move from any location on a chromosome towards another gene you want to clone but know very little about, including its sequence.
© Copyright 2001 Department of Biology, Davidson College, Davidson, NC 28036
Send comments, questions, and suggestions to: macampbell@davidson.edu