This web page was produced as an assignment for an undergraduate course at Davidson College.
Protein Analysis for GSH1 and SET4
Throughout the past two pages I have analyzed two different genes in the yeast Saccharomyces cerevisiae by looking at sequence information and microarray expression information. By doing this I discovered the vast amount of information which genomics can provide us with. In this page I will investigate these same two genes in terms of the proteins they encode and explore the developing science of proteomics. Proteomics is the study of all proteins in an organism and spans a wide variety of different methodologies.
Proteomics Tools Online
I am going to rely heavily on online tools to explore the proteomics of the two yeast genes. One of the nice resources available allows us to look at the three-dimensional structures of proteins. Visualizing a protein can be an easy way to start understanding its characteristics. The data are freely available at the RCSB Protein Data Bank. Unfortunately, they still do not have many entries in the database, and they do not have three-dimensional structures for either of my two proteins. Thus I will have to rely on other information to understand the characteristics of the proteins.
The University of Washington has a website dedicated to yeast proteomics, hosted as the NCRR Yeast Resource Center. It has data from yeast two-hybrid (Y2H) experiments, fluorescence microscopy experiments, and computer-generated protein structure predictions. In addition, the SGD has their own data, which they call "Affinity Capture-MS." I will be using these two databases.
Y2H experiments are performed to determine whether or not any two proteins interact when they are in close proximity to each other. One protein is considered to be the "bait" of the other protein, deemed the "prey." They cannot determine if the proteins actually interact in vivo, and they also do not guarantee that the proteins have their regular conformations.
The University of Washington also has data from fluorescence microscopy experiments. These allow for the visualization of where proteins are expressed. Unfortunately, neither GSH1 nor SET4 are in the data.
Researchers at Yale University performed a series of experiments on proteins in yeast using macroarrays. They used random mini-transposon insertions into the yeast as a means to determine mutant phenotypes and to observe expression levels. They then analyzed the results using a high-throughput method which resembles DNA microarrays but doesn't share some of the shortcomings DNA microarrays have. Their data is freely available at their website in what they call the TRIPLES (transposon insertion phenotypes, localization and expression in Saccharomyces) database.
One of the holy grails of proteomics is to study protein interactions at the systems-wide level. The Database of Interacting Proteins (DIP), located at the university of California at Los Angeles, is a project that focuses on graphing the interrelations between all the different proteins in organisms. This allows for the visualizations of complex networks and allows scientists to understand what proteins are doing on larger scales. GSH1 and SET4 do not have any maps available for them yet, so I cannot use this tool either at the moment.
GSH1
GSH1 is an annotated gene with well-understood functions. I therefore hope there to be lots of proteomic information available for its protein, γ-glutamylcysteine synthase. By comparing the official annotations and my previous research with the proteomic information for GSH1 I can see what proteomics has to offer that other methodologies do not have.
Y2H experiments
To begin the investigations into GSH1's proteomics, I started by looking at Y2H experimental data. There are a few different results where GSH1 was a prey protein, and several other proteins were preys. These only have 1 hit each, meaning that the results have not been confirmed by subsequent experiments.
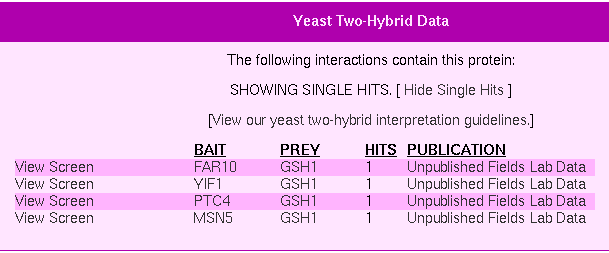
Figure 1: Y2H experimental results for GSH1 (Yeast Resource Center, 2006).
As wee see from the image, there are a total of 4 protein interactions recorded from the experiments. γ-glutamylcysteine synthase interacts with proteins from the genes FAR10, YIF1, PTC4, and MSN5. In each case, GSH1 was the prey. To understand what this means requires understanding what these four genes do. A quick click on the appropriate links tells me the following (Source: Yeast Resource Center):
- FAR10: Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far3p, Far7p, Far8p, Far9p, and Far11p; potential Cdc28p substrate.
- YIF1: Integral membrane protein required for the fusion of ER-derived COPII transport vesicles with the Golgi; interacts with Yip1p and Yos1p; localizes to the Golgi, the ER, and COPII vesicles.
- PTC4: Cytoplasmic type 2C protein phosphatase; identified as a high-copy number suppressor of the synthetic lethality of a cnb1 mpk1 double deletion; overexpression decreases high-osmolarity induced Hog1p phosphorylation and kinase activity.
- MSN5: Karyopherin involved in nuclear import and export; shown to be responsible for nuclear import of replication protein A and for export of several proteins including Swi6p, Far1p, and Pho4p; cargo dissociation involves binding to RanGTP.
Thus the Y2H experiments have given us a list of proteins with which GSH1 can interact. None of the four matches seem to be related to the understood functions for GSH1, which are free-radical and heavy metal detoxification and extra-mitochondrial protein modification. One the one hand, this means that GSH1 potentially has more functions which we are unaware of and that these interactions are a first step in discovering them. On the other hand, this might be a reflection of one the downsides of the Y2H methodology, results that are not replicated in vivo. The one trend that I do see whtihin these data is that two of the proteins, YIF1 and MSN5, are involved in cellular transport. Perhaps the interaction, even if it is artifical, is indicative of GSH1's involvment in transport mechanisms, such as movement to or from the mitochondria.
In addition to looking at the interactions between GSH1 and specific proteins, it is also possible to look at these specific proteins to see what other types of proteins they interact with. This can be a tedious task, as PTC4 alone has over 40 Y2H interactions listed.
In addition to the Y2H data from the university of Washington, the SGD has their own affinity capture data, borrowed from the BioGRID. It is surprisingly different from the data at the university of Washington. There are a total of 5 interactions, in which GSH1 is the bait 4 times and the prey once.
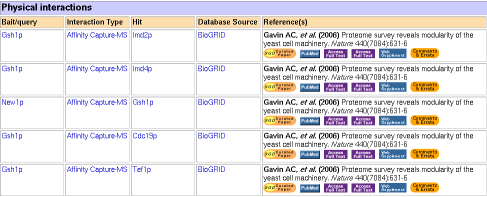
Figure 2: Physical interaction data for GSH1 (click for larger image) (Yeast Genome Database, 2006).
It turns out that none of these seem to be directly related with γ-glutamylcysteine synthesis, although perhaps they relate to the protein modification. In brief (SGD Databse):
- IMD2 and IMD4 are isozymes of inosine monophosphate dehydrogenase, which catalyze synthesis of GMP.
- CDC19 is a pyruvate kinase, converts "phosphoenolpyruvate to pyruvate, the input for aerobic (TCA cycle) or anaerobic (glucose fermentation) respiration."
- TEF1 is a translational elongation factor. It helps bind aminoacyl-tRNA to ribosomes.
- NEW1 is a binding cassette family member. It is associated with prion formation (!) and related to human Machado-Joseph Disease.
We cannot automatically assume that GSH1 does in fact interact will all these proteins. Nevertheless, it is surprising to me that none of these proteins are proteins I would expect to interact with GSH1, especially NEW1. I don't know much about Machado-Joesph Disease, but the Wikipedia article has no mentioning of prions. It would still be worthwhile to test these interactions more rigorously to see if they persist. Perhaps prion-like proteins can alter GSH1's conformation and changes its functioning. This shows that even annotated genes may not be fully understood.
Triples Data
Searching their database for GSH1 yields 8 different results. This means that there were 8 unique transposon insertion events into this gene.
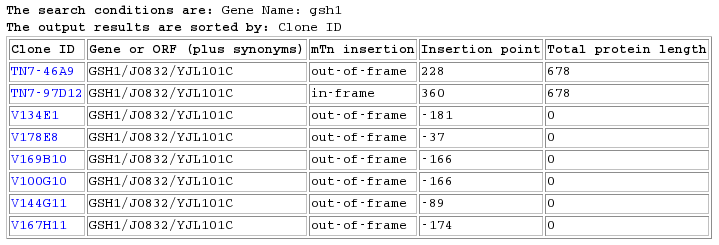
Figure 3: Database search results for GSH1 in the TRIPLES database (Triples Website, 2006).
Visiting each experimental data page, I found that in most cases the protein was expressed. The range across the difference experiments is from faint blue to blue in both the sporulative and vegetative stages. The darker the blue color the more protein is expressed. Also, one experiment, number V169B10, determined that the protein was located in the cytoplasm in both budded and unbudded stages. Only haploid cells were shown, implying that GSH1 is not vital in haploid cells but is probably vital in diploid cells.
From these data I was able to confirm that the protein is expressed in a variety of cell stages. I was also able to confirm that GSH1 is an important gene, and that many of the haploid transformants were unable to survive. The range of expression varied, possibly indicating that it is not made at constant levels. Another explanation could be that the transposon altered regulational mechanisms such as promoters, and that GSH1 actually has more consistent expression.
Database of Interacting Proteins
I was looking forward to seeing what sorts of graphs might be available for GSH1. Since it is an annotated gene and has been shown to be involved in many cellular processes, I assumed that there would be some data on GSH1. Unfortunately, there is not. Proteomics is a new field, and I expect that over time more data will accumulate in these databases.
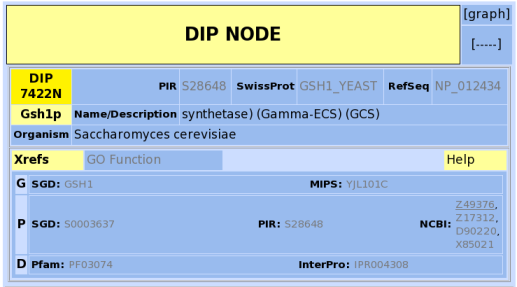
Figure 4: Screenshot of information avaible for GSH1 at the Database of Interacting Proteins (Database of Interacting Proteins, 2006).
SET4
Throughout this website I have investigated a well-annotated gene, GSH1. I have also investigated an unannotated gene with no known functions, SET4. I have previously conjectured that SET4 is a protein which interacts with histones and might methylate DNA. I also conjectured it might silence other genes. Using microarray data I analyzed when SET4 is expressed and compared it with genes it clustered with. By using proteomic tools I hope to further my conjectures and get more information about SET4.
Y2H Experiments
As described previously, Y2H experiments test for protein interactions. One protein is the bait, and the other is the prey. I searched the same databases that I used for GSH1. Below are the results:
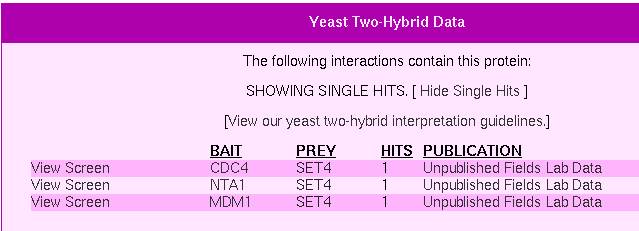
Figure 4: Y2H Experimental Results for Set4(Yeast Resource Center, 2006).
According to the yeast resource center, SET4 interacts with three other proteins as a prey. These proteins are CDC4, NTA1, and MDM1. A quick summary of what these proteins do (Yeast Resource Center, 2006):
- CDC4: F-box protein required for G1/S and G2/M transition, associates with Skp1p and Cdc53p to form a complex, SCFCdc4, which acts as ubiquitin-protein ligase directing ubiquitination of the phosphorylated CDK inhibitor Sic1p [SGD]
- NTA1: Amidase, removes the amide group from N-terminal asparagine and glutamine residues to generate proteins with N-terminal aspartate and glutamate residues that are targets of ubiquitin-mediated degradation [SGD]
- MDM1: Intermediate filament protein, required for nuclear and mitochondrial transmission to daughter buds [SGD]
One possible conclusion to draw from these interactions is that SET4 is involved in cell-cycle mechanisms. CDC4 is involved in the transitions between G1/S and G2/M.
Data from the SGD is different from the data in the YRC, and only has one interaction:
Figure 5: Physical interaction data for SET4 (click for larger image) (Yeast Genome Database, 2006).
According to the SGD, SET4 is a prey for Chl4. SGD describes it as (Yeast Genome Database):
Outer kinetochore protein required for chromosome stability, interacts with kinetochore proteins Ctf19p, Ctf3p, and Iml3p; exhibits a two-hybrid interaction with Mif2p; association with CEN DNA requires Ctf19p
The kinetochore is part of the cellular mechanism used in mitosis and meiosis. According to Wikipedia, the kinetochores consist of many proteins, including histones. This interaction provides strong reinforcement of the notion that SET4 interacts with the chromosomes and that it might bind histones. This indicates that the histones it binds might even be related to the kinetochores. Thus SET4 might play some sort of role in cell division.
Triples Data
There was a lot of useful information for GSH1 from the triples website. At first, I thought that there was no data at all for SET4. I then realized that I had to search by its official name, YJL105W/J0819, since the gene is not annotated. Searching the database yields three results:
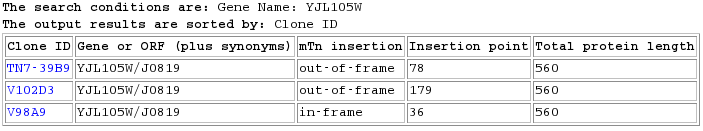
Figure 6: Database search results for YJL105W/J0819 in the TRIPLES database (Triples Website, 2006).
Looking at the data from the 3 search results, I find some things that are quite striking. First of all, in strain TN7-39B9 both the vegetative and sporulation growth conditions had intense blue LacZ expression levels. LacZ is used as a reporter gene for the protein expression, and so this indicates that SET4 is highly expressed. In fact, it is more highly expressed than GSH1 was in any of its Triples data. One possible explanation is that the yeast cells are desperately lacking SET4 and so started trying really hard to make it in large enough quantity. One way to test this would be to take this strain and inoculate it with SET4 on a plasmid - if the blue became less intense then this would confirm my hypothesis.
Strain V102D3 showed wild-type growth with an insertion at codon#179, yielding a shortened protein with regular blue LacZ expression and wild-type phenotype. This is informative because it might indicate that the shortened protein at codon #179 is still functional enough to get a regular phenotype. A quick check shows that the total number of amino acids in the full protein is 561. Thus this particular strain has a protein which is 179/561, or 31% of the full length.
The next strain, V98A9, also has a lot of data. In particular, it showed lots of different types of disrupted phenotypical data, for a total of 35. It has the insertion at codon# 36 but still most of the phenotypes are wild-type. However, the LacZ expression levels are faint and light.
Experimental Testing of My Hypotheses
Throughout this page I have mentioned a few things which I would like to test experimentally. I will explain in a little more detail what these are and how I would go about it.
There are several things which I would test for GSH1. The first one pertains to the Y2H data that indicated GSH1 might be interact with proteins involved in transportation, possibly with the Golgi apparatus. To test this, it might be possible to mutate GSH1 at different locations and then look for any differences in its location within the cell. Localization can be done using fluorescence microscopy and immunofluorescence methodologies.
Other protein interactions for GSH1 involved metabolic processes. To test for these, growing the yeast on minimal essential media and checking for different growth phenotypes could reveal whether or not GSH1 is involved in these metabolic processes. I would also be interested in investigating GSH1's possible relation to NEW1, as the idea that it might somehow be involved with prions is intriguing.
Testing other aspects of GSH1 is fun, but ultimately there is much less to learn than if you test SET4. Desiging experiments to test for SET4's functions could potentially allow for the establishment of official annotation for the gene.
A lot of data has indicated that SET4 might be cell-cycle regulated. Performing macroarray cell-cycle experiments, and looking at data for SET4, might provide information about whether or not it is regulated in this way. There is lots of cell cycle data out there from microarrays already, so using macroarrays would be interesting for comparisons. I would expect that if SET4 relates to cell divison, as indicated by its binding to a kinetochore-related protein, that it would be present during the M phase of the cell cycle in larger quantities than during other phases.
Another thing to test SET4 for would be to try to find more histone-like proteins that it might bind to. Also, it migh be possible to use Nom Dovichi's technology to see where individual SET4 protein molecules are and where they interact. I predict that it would bind histones and be near or adjacent to chromosomes. If SET4 is cell-cycle regulated and involved in the M phase, then I predict to find it in varying quantities depending on the phase, with more during M.
There is currently no solved 3d structure for SET4. One way to start analyzing its structure could be to make SET4 proteins of different lengths. I expect that longer proteins would function better than shorter proteins. Analyzing the different-length proteins in terms of activity, binding, and other characteristics could give more information about which parts of SET4 determine its ability to function.
Conclusions
GSH1 is well-annotated and well-understood. By using proteomics tools I was able to gather more information about it. Most of the information confirms my previous findings. Some of the information, however, lends itself to the possibility that GSH1 does more than we know. I suggested a few ways to test whether or not this is true.
The proteomics data for SET4 has proved itself to be quite intriguing. There are a lot of Y2H-type protein interactions that have been discovered in the lab. Some of these interactions are with proteins related to mechanisms in cell division. Thus the data supports the notion that SET4 is located within the nucleus and somehow interacts with chromosomes. Triples data confirms that the is expressed and shows that there are a lot of different mutant phenotypes. By compiling all this information, I was able to develop several possible experiments to test whether or not my predictions for SET4's functions are correct.
References
Online Abstracts
Molin, M. and A. Blomgerg. Dihydroxyacetone detoxification in Saccharomyces cerevisiae involves formaldehyde dissimilation [abstract]. <http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=search&term=gsh1+proteomics> Mol Microbiol. 2006 May;60(4):925-38.
Online References
CDC19/YAL038W Summary. Yeast Genome Database. 2006. <http://db.yeastgenome.org/cgi-bin/locus.pl?locus=cdc19> Accessed 2006 Nov 17.
CHL4/YDR254W Summary. Yeast Genome Database. 2006. <http://db.yeastgenome.org/cgi-bin/locus.pl?locus=YDR254W> Accessed 2006 Nov 17.
GSH1/YJL101C Physical and Genetic Interactions. Yeast Genome Database. 2006. < http://db.yeastgenome.org/cgi-bin/phenotype/phenotype.pl?dbid=S000003637&type=allinteractions> Accessed 2006 Nov 16.
IMD2/YHR216W Summary. Yeast Genome Database. 2006. <http://db.yeastgenome.org/cgi-bin/locus.pl?locus=imd2> Accessed 2006 Nov 17.
IMD4/YML056C Summary. Yeast Genome Database. 2006. <http://db.yeastgenome.org/cgi-bin/locus.pl?locus=imd4> Accessed 2006 Nov 17.
Kinetochore. Wikipedia, 2006. <http://en.wikipedia.org/wiki/Kinetochore> Accessed 2006 Nov 17.
Machado-Joseph Disease. Wikipedia, 2006. <http://en.wikipedia.org/wiki/Machado-Joseph_Disease>Accessed 2006 Nov 17.
NEW1/YPL226W Summary. Yeast Genome Database. 2006. <http://db.yeastgenome.org/cgi-bin/locus.pl?locus=new1> Accessed 2006 Nov 17.
TEF1/YPR080W Summary. Yeast Genome Database. 2006. <http://db.yeastgenome.org/cgi-bin/locus.pl?locus=tef1> Accessed 2006 Nov 17.
Triples Website. 2006. <http://ygac.med.yale.edu/triples/default.htm> Accessed 2006 Nov 16.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=531502> Accessed 2006 Nov 9.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=528729> Accessed 2006 Nov 9.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=529694> Accessed 2006 Nov 9.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=532431> Accessed 2006 Nov 9.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=533417> Accessed 2006 Nov 9.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=531506&showDescriptions=false&showSingles=true> Accessed 2006 Nov 16.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=530206> Accessed 2006 Nov 16.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=531688> Accessed 2006 Nov 16.
YRC Public Data Repository - View Protein Information. 2006. <http://www.yeastrc.org/pdr/viewProtein.do?id=532779> Accessed 2006 Nov 16.