This
web page was produced as an assignment for an undergraduate course at
Davidson College.
Assignment
1
What
was the research project?
Researchers
were interested to figure out how free living chitrid fungi infect
their amphibian hosts, so they sequenced the genomes of both Batrachochytrium
dendrobatidis (Bd), and Batrachochytrium salamandrivorans (Bsal),
in conjunction with the genomes of two other free-living, fungal
pathogens to ascertain which genes allowed Bd and Bsal to infect and
cause disease in amphibians.
Were
researchers testing a hypothesis or doing discovery science?
The
researchers were doing discovery science, as they were trying to
discover which genes allowed the infection and caused disease in
amphibians.
What
genomic technology was used in this project?
Genomic
sequencing using paired-end reads and Sanger technology.
What
is the take home message?
The
take home message of this study is that, while Bd and Bsal are
related, and both affect amphibians, each fungus has specific
gene-family traits that cause each virus to have distinct infection
strategies. Furthermore, it is important to continue studying
the genetic differences in chytrid funguses in order to understand how
they are likely to affect amphibians in the future, so protections can
be put into place that would preserve biodiversity.
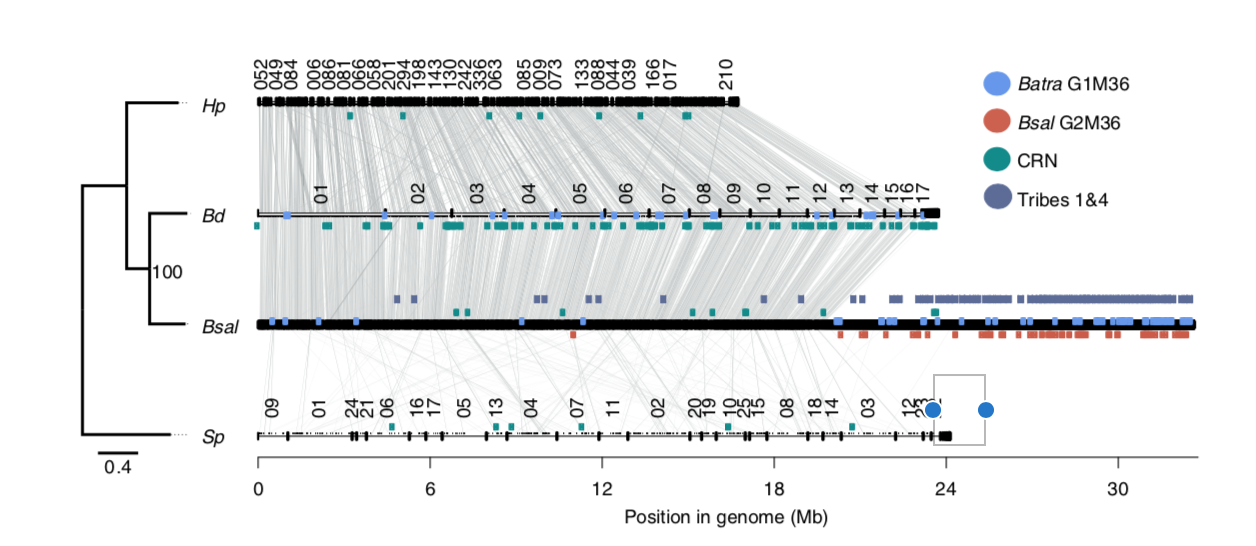
Figure
1: Phylogenetic tree showing
relationships between four different chytrid funguses (Farrer et
al. 2017).
My
evaluation of the project:
The
discovery of Bsal, and its consequent effect on salamanders from the
pet trade make it extremely important for scientists to know how the
fungus spreads to and infects its host, so that precautions can be
taken to protect the amphibian biodiversity in regions where the
fungus has not yet spread. The study is particularly valuable
because not only did researchers compare fungus genomes, they also
compared the effects of each fungus on salamander (T. wenxianensis)
skin. The news article reporting on this study is good in that
it delivers the take away message clearly, and gives good insight into
why this study was important. However, I think the article could
have done a better job accurately reporting how the data was gathered
in this study. Specifically, the news article makes no mention
of the two other funguses that were analyzed, and gives some
misleading information about the methods of the study, in particular
stating that only one live salamander was used in the study, when
there were nine salamanders used.
Literature
Cited:
Brogan,
C. "Breakthrough in 'amphibian plague': Deadly fungus genes
identified." Phys.org. 27 March 2017. Web. 4 February 2018.
Farrer,
R., Martel A., Verbrugghe, E., Abouelleil, A., Ducatelle, R.,
Longcore, J., James, T., Pasmans, F., Fisher, M. and Cuomo, C.
"Genomic innovations linked to infection strategies across
emerging pathogenic chytrid fungi." Nature Communications. 8:14742
(2017).