This web page was produced as an assignment for an undergraduate course at Davidson College.
My Favorite Yeast Proteins:
SPR3 and YGR058W
SPR3 and YGR058W are genes within the Saccharomyces cerivisiae species. They are located on chromosome VII within yeast. SPR3 is an annotated gene whose product is a septin protein within spore walls that is involved with sporulation, cellular mophogenesis, and spore wall assembly. YGR058W is a non-annotated ORF, thus meaning much of its protein function has not been documented. Using various genomics databases I have found large amounts of information about these two genes, primarily involving each gene's nucleotide and protein sequences and their expression patterns in certain experimental conditions. These databases have helped me to confirm SPR3's functions and allowed me to hypothesize the functions of YGR058W.
In this assignment, I will use databases specifically related to proteomics to learn more about the functions of the proteins SPR3 and YGR058W. This analysis of these proteins will hopefully illuminate the characteristics of these proteins and interactions it may have within a cell.
________________________________________________________________________________
SPR3 (YGR059W)
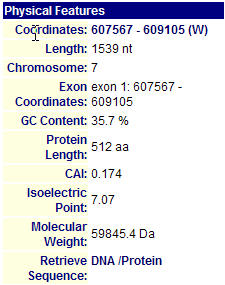
I first searched the MIPS database to find information on the SPR3 protein. This database gave me a good deal of useful information on the physical features of this protein. As seen below in Figure 1, the protein has a length of 512 amino acids, an isoelectric point of 7.07, and a molecular weight of 59845.4. A protein's isoelectric point is the pH of its local environment.

Figure 1. MIPS Physical Features Information of SPR3. One should note that the protein is 512 amino acids long, has a Isoelectric point of 7.07, and a molecular weight of 59845.4 Da. (Image from DIP, 2004: http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YGR059w&db=CYGD)
From the same database, I looked at proteins that had been found to interact with SPR3. The results are below in Figure 2. Unfortunately the link(seen below in blue) was inactive, so I couldn't see the nature of the interactions with SPR3. KAR5, MYO1, and SPR28 are all shown to have protein-protein interactions with SPR3. A quick search at SGD showed that: KAR5 is located in the membrane of the endoplasmic-reticulum and involved in conjugation in cell fusion, MYO1 is a microfilament involved in cytokinesis, and that SPR28 is a septin-related protein that is located in the cytoskeleton, is involved in sporulation, cellular wall organization and genesis, and cellular morphogenesis just as SPR3 is.
![]()
Figure 2: Protein Interactions with SPR3. KAR5, MYO1, and SPR28 are shown to interact with SPR3. The (htp) next to KAR5 and MYO1 indicate that the interactions were apparent only in high throughput experiments. (Image from DIP, 2004: http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YGR059w&db=CYGD)
Next I used the Database of Interacting Proteins(DIP) to see other proteins that interact with SPR3. In Figure 3 you can see that SPR3 is known to interact with 10 other proteins. The most directly interactive proteins(represented by the orange nodes) are MYO1 at the top of the figure and KAR5 at the bottom. The second tier of interacting proteins which are less directly interative with SPR3 consist of SMC2, SMC3 at the bottom and MLC2, MYO4, MLC1, MYO2, ACT1 and SRO7 at the top. The direct interactions with KAR5 and MYO1 are consistent with the protein interactions from MIPS in Figure 2.
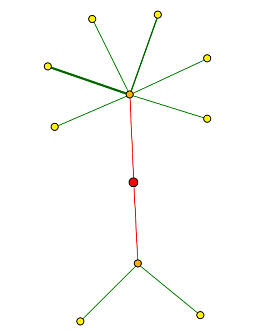
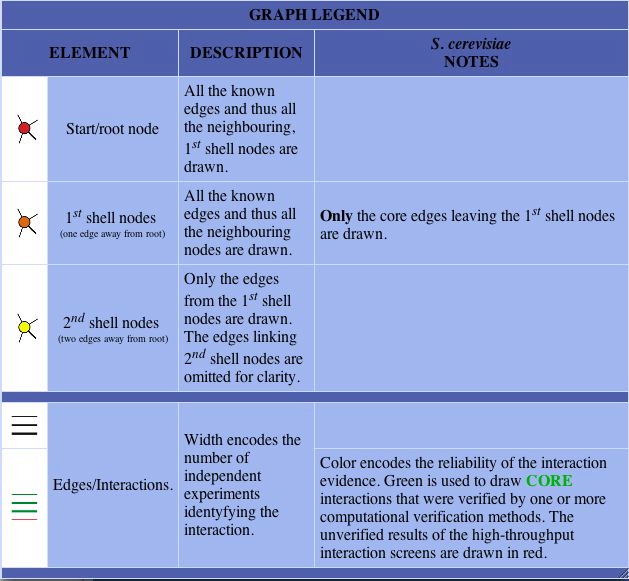
Figure 3. 10 Proteins known to interact with SPR3. The red node represents SPR3. The orange nodes represent the proteins known to directly interact with SPR3. The yellow nodes represent a second tier of proteins that interact more indirectly than the orange nodes. The lines are used to indicate the interactions and the thicker green line indicates a greater reliability that the proteins interact (as the legend indicates). (Image from DIP,2004: http://dip.doe-mbi.ucla.edu/dip/DIPgraph.cgi?PK=3011&D=2).
I then searched the Benno Figure 1 PDF for SPR3. The figure is from a paper that showed protein interactions by Stan Fields, Peter Uetz, and Benno Schwikowski. The file shows many protein-protein interactions as these authors determined. Figure 3 shows SPR3 only interacts with 1 other protein directly, SPR28. SPR3 is linked to SPR28 by a black line which means that they have different cellular roles and localizations. The yellow boxes indicate unknown roles for the proteins.
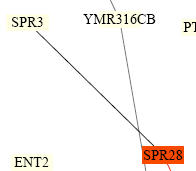
Figure 4. SPR3 interactions from "Benno Figure 1" by Fields, Uetz, and Schwikowski. The proteins are shown in boxes and lines indicate interactions known to occur between the two proteins. SPR3 is seen to interact with just one protein, SPR28. The black lines mean the proteins have different cellular roles and different localizations and the yellow boxes mean unknown roles of the proteins.(Image from http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/seq/Benno/NB_Figure1color.pdf)
I then searched the Pathcalling Yeast Interactive Database. The results for SPR3 can be seen below in Figure 5. The proteins that are seen to interact with SPR3 are SPR28 and CDC11. The result of SPR28 is concurrent with the interaction seen above in the Benno Figure 1. According to this database, CDC11 is involved in proper bud growth during sporulation which makes it similar to SPR3 in that each has a role in sporulation.
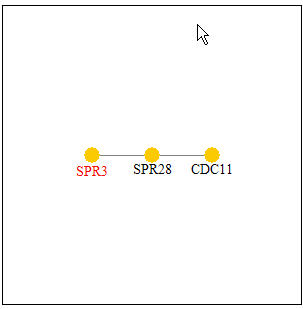
Figure 5: SPR3's Protein Interactions. SPR3(located on the left) is shown to interact with SPR28 directly and indirectly with CDC11.(Image from Pathcalling, 2004: http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastGene?geneIdIn=2253&keywordIn=spr3).
Figure 2-5 show that though SPR3 was seen in the MIPS and DIP database to interact with the KAR5 and MYO1 proteins, it was difficult to determine the role of SPR3 that allowed this to happen as these two proteins had starkly different protein functions than SPR3. Thus, to further confirm SPR3's known protein functions I found that most of the databases I used showed an interaction between SPR28 and SPR3. Because SPR28 encodes a septin-related protein and has nearly identical protein functions I chose to focus on this protein to show how protein-protein interactions can be used to make statements about each proteins function. Obviously deciphering a protein's functions is difficult despite the wealth of information available about the interactions among proteins. Sometimes these interactions imply that these proteins have similar functions and sometimes these interactions are between two proteins that come into contact despite being involved in different functions altogether, or functions that neither was known to be involved with before. |
There were multiple databases that either provided no information at all about the SPR3 protein or whose information was not helpful in determining the function of the protein. Please note each case below:
TRIPLES: 1 result appeared for this database, but the information was not useful for determining the function of the protein. (http://ygac.med.yale.edu/triples/get_clone_info.asp?cloneid=TN7-13H4)
Yeast-2-Hybrid: No results
Interaction Sequence Tags: No results
Enzymes and Metabolic Pathways: No results
PDB: No results. Also no results for PDB homologues at SGD's site.
ExPaSy: There were two results, but both of them consisted of Swiss-PROT information that was involved sequence analysis and wasn't too helpful in helping to determine the function of the protein.
Yeast Grid: No results for SPR3. Results shown here.
Benno Figure 2: No results for SPR3.
Degradation, Aging, and Membrane PDF files: SPR3 is present but little information about the protein can be obtained other than that it interacts with SPR28.
____________________________________
Final Thoughts and New Experiments
In three webpages I have now analyzed the sequence, expression patterns, and protein interactions and functions of the annotated gene SPR3 using online genomic databases. These databases indicate that SPR3 is a septin protein, which has a molecular functions of being a structural component of the cytoskeleton, whose biological processes include cell wall organization, cellular morphogenesis, and spore wall assembly during sporulation, and whose cellular component is within the prospore membrane, septin ring, and spore wall. These different databases allowed me to compare genes and proteins with similar sequences, transcription patterns under certain conditions, and protein function. I only found two proteins which consistently interacted with SPR3, (SPR28 is a septin-related protein involved in sporulation and CDC11 is also known to be involved in sporulation) and both of these had very similar protein functions. The proteomics databases provided a substantial amount of information, alot of which was redundant, however, there were also a large amount of databases that had no information for this annotated gene. Hopefully, in tiime more of these databases will provide for information about SPR3 and other yeast genes to further confirm this protein's - and other proteins' - functions.
As for for new experiments, I suppose I would design experiments that could further test SPR3's role in the cell. For one I think an immunostaining experiment that tagged SPR3 with a fluorescent glowing antibody would further confirm it's location in yeast cells: outside of the nucleus and in cytoskeleton structures.
_______________________________________________________________________________
YGR058W
In the first web page I found out that the cellular component was known for this ORF, as it was found in the nucleus and cytoplasm, however it's molecular function and biological process was unknown. In my last web page, I hypothesized that the unannotated ORF YGR058W's molecular function is protein kinase activity and its biological process was involvement in iron ion homeostasis or transport.
First I searched the MIPS database for the YGR058W protein.
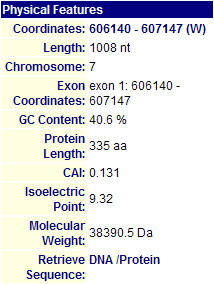
Figure 1. MIPS Physical Features Information of YGR058W. This protein has 335 amino acids. It has an isoelectric point of 9.32, and a molecular weight of 38390.5 Da. (Image from MIPS, 2004: http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YGR058w&db=CYGD)
From the MIPs database, I found proteins found to interact with YGR058W. The results are below in Figure 2. Yet again the link was inactive, so I couldn't see the nature of the interactions with SPR3. The proteins found to interact with this ORF were YNL047C, YOR264w, YDL203c, YOR097c, HOG1, LSB1, and TEM1.

Figure 2: Protein Interactions with YGR058W. YNL047C, YOR264w, YDL203c, YOR097c, HOG1, LSB1, and TEM1 are shown to interact with YGR058W. (Image from MIPS, 2004: http://mips.gsf.de/genre/proj/yeast/searchEntryAction.do?text=YGR058w&db=CYGD)
I then searched DIP to find information on the proteins with which YGR058W interacts. The results were 50 interacting proteins, which is substantially larger than the amount I found for SPR3. This is pretty surprising to me since YGR058W is an ORF and its function is not fully known. The proteins most directly interactive(in orange nodes) with YGR058W are YDL203C, YOR097C, HOG1P, TEM1P, LSB1P, DSE3P, and YNL047C.
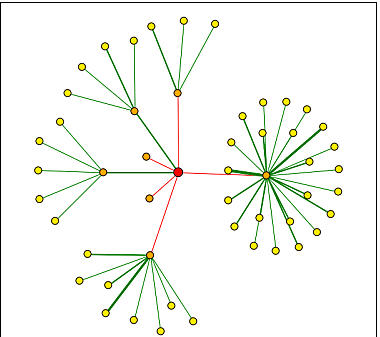
Figure 3. YGR058W interactions with other proteins. YGR058W interacts with 50 other proteins. The yellow nodes represent a second tier of proteins that interact more indirectly than the orange nodes. The lines are used to indicate the interactions and the thicker green line indicates a greater reliability that the proteins interact(See legend in Figure 3 for SPR3 for further details).(Image from DIP, 2004: http://dip.doe-mbi.ucla.edu/dip/DIPgraph.cgi?PK=1862&D=2)
I then searched within the Benno Figure 1 to see the interactions that were determined by this experiment. Though it is pretty hard to decipher from the picture, the proteins that interact with YGR058W are HOG1, YOR264W, YGR136W, and YDL203C.
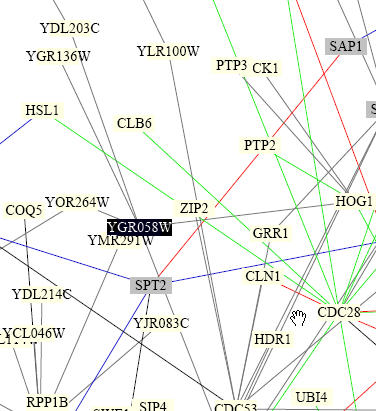
Figure 4. YGR058W interactions from "Benno Figure 1" by Fields, Uetz, and Schwikowski. The proteins are shown in boxes and lines indicate interactions known to occur between the two proteins. YGR058W is seen to interact with four proteins: HOG1, YOR264W, YGR136W, and YDL203C .(Image from http://occawlonline.pearsoned.com/bookbind/pubbooks/bc_mcampbell_genomics_1/medialib/seq/Benno/NB_Figure1color.pdf).
I then searched the Pathcalling Yeast Interactive Database for YGR058W. The results for SPR3 can be seen below in Figure 5. The proteins that are seen to directly interact with SPR3 are YNL047C, HOG1, YGR136W, YDL203C, and YOR264W. All of these results except YNL047C concur with the interactions seen above in the Benno Figure 1. YNL047C, HOG1, and YDL203 concur with Figure 2 from DIP.The only protein that is annotated is HOG1 and it is a mitogen-activated protein kinase(MAP kinase) that is involved in osmoregulation.
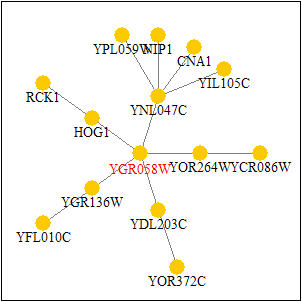
Figure 5: YGR058W's Protein Interactions. The nodes indicate proteins and the lines indicate interactions. As one see HOG1, YNL047C, YOR264W, YDL203C, and YGR136. (Image from Pathcalling, 2004: http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastGene?geneIdIn=2252&keywordIn=ygr058w)
I then searched Yeast Grid. It shows seven proteins known to have interactions with YGR058W. These interactions were determined by the Yeast Two Hybrid method which a bait protein draws prey protein, thus confirming an interaction.
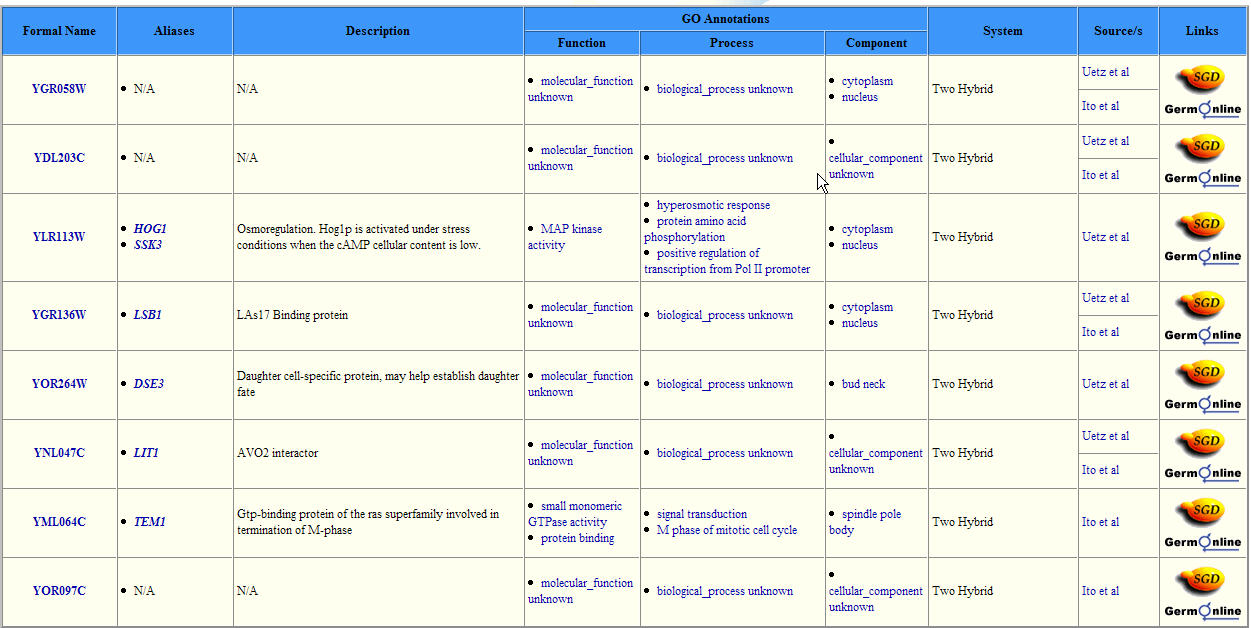 Figure 6: YGR058W's Protein Interactions as determined by Y2H method. This grid show proteins found to interact with the YGR058W protein. (Image from SGD, 2004: http://biodata.mshri.on.ca/yeast_grid/servlet/SearchResults?keywords=YGR058W)
Figure 6: YGR058W's Protein Interactions as determined by Y2H method. This grid show proteins found to interact with the YGR058W protein. (Image from SGD, 2004: http://biodata.mshri.on.ca/yeast_grid/servlet/SearchResults?keywords=YGR058W)
Figures 2 through 6 represent data from multiple proteomics databases that indicate interactions between the unannotated ORF YGR058W and other proteins. The most consistently interactive annotated gene is HOG1. This interaction implies some similarity in biological process and molecular function, so because HOG1 is a MAP-kinase involved in the regulation of water and other minerals within internal fluids such as blood, I am going to infer that YGR058W's protein functions consist of those of HOG1. YGR058W is also seen to interact with multiple other proteins, however these are unannotated so not particularly helpful in illuminating the protein function of YGR058W. However, if these proteins' functions are ever determined, they would prove very helpful in determining the functions of YGR058W. |
There were multiple databases that either provided no information at all about the YGR058W protein or whose information was not helpful in determining the function of the protein. Please note each case below:
TRIPLES: No results for YGR058W
Yeast-2-Hybrid: No results for YGR058W
Interaction Sequence Tags: No results for YGR058W
Enzymes and Metabolic Pathways: No results for YGR058W
PDB: No results.
ExPaSy: Just as for SPR3 there were two results, but both of them consisted of Swiss-PROT information that was involved sequence analysis and wasn't too helpful in helping to determine the function of the protein.
Benno Figure 2: No results for YGR058W
Y2H: This database indicates interactions between YOR264W and HOG1, but this was already known.
____________________________________
Final Thoughts and New Experiments
I have now analyzed the unannotated ORF , YGR058W, and its sequence, expression patterns, protein functions and interactions. In previous web assignments I had found that it's cellular component of being in the nucleus and cytoplasm was known, however both its molecular function and biological process were unknown. In the two web assignments linked above I hypothesized that YGR058W's molecular function was protein kinase activity and the biological process was being involved in iron ion homeostasis or transport. From this most recent analysis, that of using proteomics, I think that the interaction with HOG1 is the most telling clue of YGR058W's protein function, and because HOG1 is a is a mitogen-activated protein kinase(MAP kinase) that is involved in osmoregulation. This is consistent with the molecular function I hypothesized in the previous webpage is further legitimized. Because of these expression pattern and proteomics information I hypothesize that YGR058W acts as some kind of protein kinase, and whose biological process is most likely osmoregulation, as taken from the interaction with HOG1. Osmoregulation is the control of the levels of water and mineral salts in the blood. Though these protein functions are still highly speculative, this is also consistent with my previous hypothesis of YGR058W's involvement in iron-ion transport, as iron is an integral component of blood.
One idea for new experiments to determine more about YGR058W would be immunostaining for YGR058W. It would be interesting to see it's location as its cellular component is known to be in the nucleus and the cytoplasm. On a bigger scale if it was expressed predominantly in blood cells or cells of other internal fluids, as this would concur with its role in the osmoregulation process.
Another experiment would be to mutate the YGR058W in some way(using knockout or other method), to see the phenotype of yeast when insufficient YGR058W protein was present. This would at the bare minimum indicate YGR058W's importance to the survival of the yeast organism - if it caused death or not. If the organism did survive it would be interesting to see if any abnormalities were seen, and in what areas and processes these abnormalities occurred; these abnormalities should allow one to determine further the role of the YGR058W protein with the cell and overall within the organism.
References
[DIP] Database of Interacting Proteins. 2004. <http://dip.doe-mbi.ucla.edu/dip/Search.cgi?SM=3>. Accessed 2004 Nov. 18.
MIPS Comprehensive Yeast Genome Database. 2004. <http://mips.gsf.de/genre/proj/yeast/index.jsp>. Accessed 2004 Nov. 19.
PathCalling Yeast Interaction Database. 2004. <http://portal.curagen.com/extpc/com.curagen.portal.servlet.PortalYeastList?modeIn=List>. Accessed 2004 Nov. 18.
[SGD] Saccharomyces Genome Database. 2004. <http://www.yeastgenome.org/>. Accessed 2004 Nov. 19.
Yeast Grid. 2004. < http://biodata.mshri.on.ca/yeast_grid/servlet/SearchPage> Accessed 2004 Nov. 19.
________________________________________________________________________________
Questions, Comments? E-mail John